Identifying compound classes through machine learning

Everything that lives has metabolites, produces metabolites and consumes metabolites. These molecules arise as intermediate and end products from chemical processes in an organism's metabolism. Therefore, they not only have huge significance for our lives, but they also provide valuable information about the condition of a living being or an environment. For example, metabolites can be used to detect diseases or, in the field of environmental technology, to examine drinking water samples. However, the diversity of these chemical compounds causes difficulties in scientific research. To date, only few molecules and their properties are known. If a sample is analyzed in the laboratory, only a relatively small proportion of it can be identified, while the majority of molecules remain unknown.
Bioinformaticians at Friedrich Schiller University Jena, Germany together with colleagues from Finland and the USA, have now developed a unique method with which all metabolites in a sample can be taken into account, thus considerably increasing the knowledge gained from examining such molecules. The team reports on its successful research in the renowned scientific journal Nature Biotechnology.
Learning, recognizing and assigning structural properties
"Mass spectrometry, one of the most widely used experimental methods for analyzing metabolites, identifies only those molecules that can be uniquely assigned by matching them against a database. All other, previously unknown, molecules contained in the sample do not provide much information," explains Prof. Sebastian Böcker from the University of Jena. "With our newly developed method, called CANOPUS, however, we also obtain valuable insight from the unidentified metabolites in a sample, as we can assign them to existing compound classes."
CANOPUS works in two phases: first, the method generates a 'molecular fingerprint' from the fragmentation spectrum measured by means of mass spectrometry. This contains information about the structural properties of the measured molecule. In the second phase, the method uses the molecular fingerprint to assign the metabolite to a specific compound class without having to identify it.
Learning from the data
"Machine learning methods usually require large amounts of data in order to be trained. In contrast, our two-stage process makes it possible in the first step to train on a comparatively small amount of data of tens of thousands of fragmentation mass spectra. Then, in the second step, the characteristic structural properties that are significant for a compound class can be determined from millions of structures," explains Dr. Kai Dührkop from the University of Jena.
The system therefore identifies these structural properties in an unknown molecule within a sample and then assigns it to a specific compound class. "This information alone is sufficient to answer many important questions," Böcker emphasizes. "The precise identification of a metabolite would be far more complex and is often not necessary at all." The CANOPUS method uses a deep neural network predicting around 2,500 compound classes.
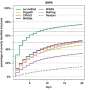
With their method, the Jena bioinformaticians have compared, for example, the intestinal flora of mice in which one experimental group had been treated with antibiotics. The examinations show which metabolites the mouse and its intestinal flora produce. Such research results can provide important information about the human digestive and metabolic system. Through two further application examples, which they present in their study, the Jena scientists demonstrate the functionality and power of the CANOPUS method.
Jena molecule search engine used millions of times
With the new method, the bioinformaticians from Jena are expanding the possibilities of the search engine for molecular structures "CSI:FingerID", which they have been making available to the international research community for around five years. Researchers around the world now use this service thousands of times a day to compare a mass spectrum from a sample with various online databases, in order to identify a metabolite more precisely. "We are approaching the one hundred millionth request and we are sure that CANOPUS will further increase the number of users," says Sebastian Böcker.
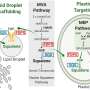
The new process strengthens the field of metabolomics—that is, research on these omnipresent small molecules—and increases its potential in many research areas, such as pharmaceuticals. Many active pharmaceutical substances in use for decades, such as penicillin, are metabolites; others could be developed with their help.
More information: Dührkop, K., Nothias, LF., Fleischauer, M. et al. Systematic classification of unknown metabolites using high-resolution fragmentation mass spectra. Nat Biotechnol (2020). doi.org/10.1038/s41587-020-0740-8
Journal information: Nature Biotechnology
Provided by Friedrich Schiller University of Jena