Viral dark matter exposed: Metagenome database detects phage-derived antibacterial enzyme
In a pioneer study published in Cell Host & Microbe, Researchers at Osaka City University and The Institute for Medical Science, The University of Tokyo, reported intestinal bacterial and viral metagenome information from the fecal samples of 101 healthy Japanese individuals. This analysis, leveraging host bacteria-phage associations, detected phage-derived antibacterial enzymes that control pathobionts. As proof of concept, phage-derived endolysins are shown to regulate C. difficile infection in mice.
Abnormalities in human intestinal microflora, known as dysbiosis, are connected to various diseases. Altered microbial diversity impairs the beneficial effects of host intestinal microflora, which cause some symbiotic commensal bacteria to acquire virulence traits, proliferate, and become directly involved in the development of disease. These bacteria are referred to as "pathobionts," which are distinct from opportunistic pathogens.
C. difficile, which is a Gram-positive, spore-forming anaerobic bacterium, is a pathobiont and the representative cause of nosocomial diarrhea following antibiotic treatment. Since antibiotic usage has the risk of killing beneficial bacteria and promoting dysbiosis, the development of methods to specifically manipulate intestinal pathobionts is essential.
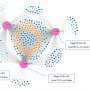
"Phages were sure to be applicable as a highly specific therapy for intestinal pathobiont elimination," believed Professor Satoshi Uematsu. The infectious associations between phages and bacteria in the human intestine is essential information for the development of phage therapies. Known as "viral dark matter" as it had yet to be understood, researchers obtained metagenome information about bacteria-phage associations from the fecal samples of 101 healthy individuals through the development of a virome analysis pipeline. Based on this information, researchers screened C. difficile-specific phages and identified novel antibacterial enzymes, both in vitro and in vivo.
"The accumulation of more metagenomic information on intestinal phages and bacteria will open up the possibility of developing treatments for a variety of dysbiosis-related diseases," say Dr. Kosuke Fujimoto and Prof. Seiya Imoto.
More information: Cell Host & Microbe (2020). DOI: 10.1016/j.chom.2020.06.005
Journal information: Cell Host & Microbe
Provided by Osaka City University