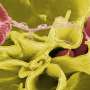
Bacteria that are persistently resistant to one antibiotic are 'primed' to become multidrug-resistant bugs

Antibiotics save lives—but using them also helps antibiotic-resistant strains evolve and spread. Each year, antibiotic-resistant bacteria infect some 2.8 million people in the United States, killing more than 35,000, according to the Centers for Disease Control and Prevention. Infections by multidrug-resistant—or MDR—bacteria, which are resistant to two or more antibiotics, are particularly difficult to treat.
Scientists at the University of Washington and the University of Idaho have discovered just how readily MDR bacteria can emerge. In a paper published April 6 in Nature Ecology & Evolution, the researchers report that, for a bacterial pathogen already resistant to an antibiotic, prolonged exposure to that antibiotic not only boosted its ability to retain its resistance gene, but also made the pathogen more readily pick up and maintain resistance to a second antibiotic and become a MDR strain.
The team's experiments indicate that prolonged exposure to one type of antibiotic essentially "primed" the bacteria. This priming effect made it more likely that the bacteria would acquire resistance to additional antibiotics, even in the absence of further antibiotic exposure, and helped the strain hold on to those antibiotic-resistance traits for generations.
"Exposure to antibiotics appears to select indirectly for more stable antibiotic resistance systems," said Benjamin Kerr, a UW professor of biology and co-senior author on the paper. "A more stable system in a strain will increase the chances that it will acquire resistance to multiple antibiotics."
Their findings also show how antibiotic exposure affects the evolutionary dynamics within bacteria.
"This could help explain not only the rise of multidrug resistance in bacteria, but also how antibiotic resistance persists and spreads in the environment—in health care settings, in soil from agricultural runoff—even long after the antibiotic exposure has ended," said co-senior author Eva Top, a professor of biology at the University of Idaho.
The researchers tested a common mechanism for the spread of antibiotic resistance: plasmids. These are circular strands of DNA that can contain many types of genes, including genes for antibiotic resistance. Bacteria easily share plasmids, even across species.
Yet plasmids have their downsides, and past research has shown that bacteria readily shed them.
"Even though they can carry beneficial genes, plasmids can also interfere with many types of processes inside a bacterial cell, such as metabolism or DNA replication," said lead author Hannah Jordt, a UW research scientist in biology. "So, scientists have generally thought of plasmids as costly and burdensome to the host cell."
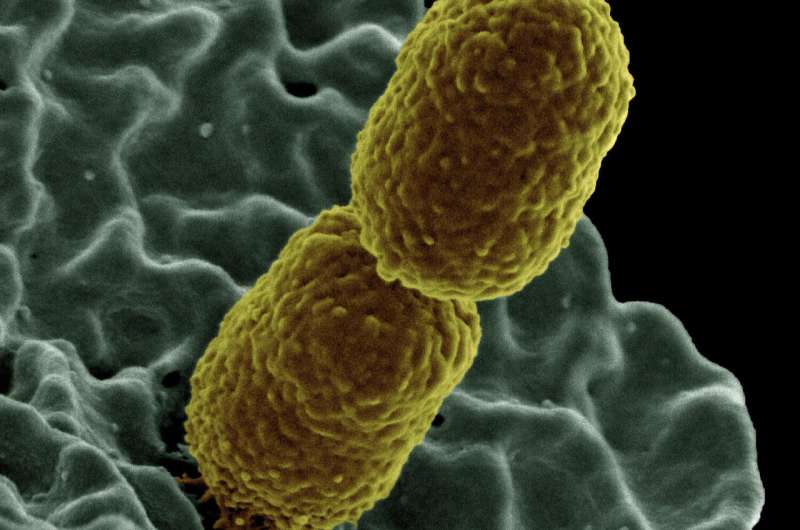
The UW-University of Idaho team worked with E. coli cells containing a tetracycline-resistance plasmid and Klebsiella pneumoniae cells containing a chloramphenicol-resistance plasmid. Both hosts, which had not been grown in the presence of antibiotics before, showed no great loyalty to their plasmids. After nine days in an antibiotic-free environment, the fraction of Klebsiella still harboring a plasmid dropped to less than 50%. For E. coli, less than 20% kept their plasmid.
When the researchers exposed the strains to antibiotics, growing each for 400 generations in their respective antibiotic, the strains showed greater affinity for their plasmids even after the antibiotic threat was lifted. After nine days in an antibiotic-free growth medium, more than half of E. coli and Klebsiella cells held on to their respective plasmid.
"Of course, the cells needed their plasmids to help them survive the antibiotic exposure. But even after we took away that selective pressure, both strains retained their plasmids at significantly higher levels than they had before the antibiotic exposure," said Jordt.
In addition, other experiments showed that antibiotic exposure increased the occurrence of MDR Klebsiella. Even without exposure to antibiotics, Klebsiella pneumoniae can acquire multiple plasmids. For example, when the researchers grew antibiotic-naive Klebsiella and E. coli plasmid-bearing strains together, a small fraction of Klebsiella became MDR by retaining their chloramphenicol-resistance plasmid and acquiring tetracycline-resistance plasmids from E. coli. But when the researchers repeated the experiment using antibiotic-exposed bacteria, they found roughly 1,000 times more MDR Klebsiella.
Prior prolonged exposure to just one antibiotic—chloramphenicol—had increased the likelihood that chloramphenicol-resistant Klebsiella would acquire the tetracycline-resistance plasmid from E. coli in an antibiotic-free environment. In addition, the team's experiments also showed that, when MDR cells were grown later in an antibiotic-free environment, chloramphenicol-exposed Klebsiella more readily held on to both resistance plasmids.
Evolution can explain both the persistence of the antibiotic-resistance plasmids and the increase of MDR in Klebsiella, the researchers say: Exposing the strains to their respective antibiotic selected for mutations in their genomes to minimize the clash between plasmid and host, making it less costly to keep that plasmid as well as others.
"We believe that by stabilizing one plasmid, these mutations make them more likely to stabilize additionally acquired plasmids," said Kerr.
Additional experiments may identify the specific mutations that helped E. coli and Klebsiella keep their plasmids, and why Klebsiella was more able to develop into a MDR strain than E. coli. Even though antibiotic exposure creates selective pressure to keep plasmids and acquire new ones, there is still widespread variation in how different species keep, share and receive plasmids.
"There are many, many details to be worked out in the future," said Top. "But what we see here is that even short-term exposure to just one antibiotic accelerates the development of multidrug resistance, which should give us pause as we use these drugs in health care, agriculture and other settings."
More information: Hannah Jordt et al, Coevolution of host–plasmid pairs facilitates the emergence of novel multidrug resistance, Nature Ecology & Evolution (2020). DOI: 10.1038/s41559-020-1170-1
Journal information: Nature Ecology & Evolution
Provided by University of Washington