Novel synthetic proteins rival their natural counterparts in proton transport
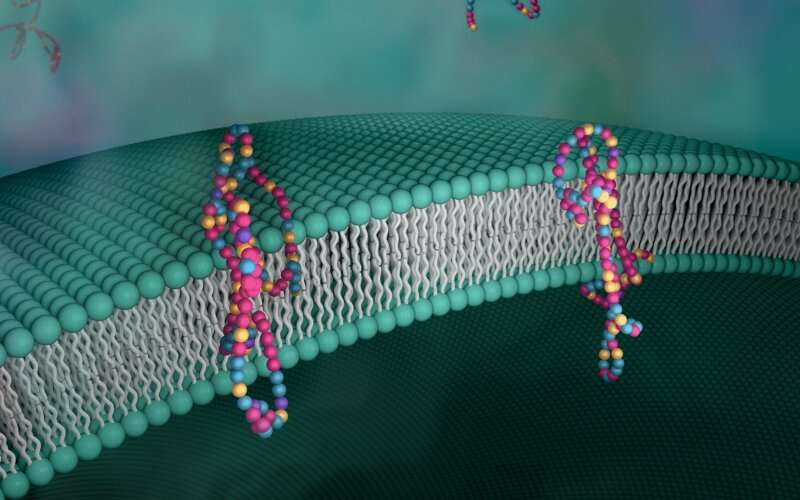

Biological membranes, such as the "walls" of most types of living cells, primarily consist of a double layer of lipids, or "lipid bilayer," that forms the structure, and a variety of embedded and attached proteins with highly specialized functions, including proteins that rapidly and selectively transport ions and molecules in and out of the cell.
Artificial membranes have been used for small- and large-scale industrial processes since the middle of the twentieth century, yet their inefficiency can make some processes relatively slow and expensive. Scientists have long sought to develop synthetic membranes that could match the selectivity and high-speed transport offered by their natural counterparts.
Now a team led by University of California Berkeley researchers has designed and—using neutron scattering at Oak Ridge National Laboratory (ORNL)—also accurately characterized a novel polymer that is as effective as natural proteins in transporting protons through a membrane. The results of their research were published in Nature.
This major milestone has the potential to transform a wide range of technologies, such as making batteries and water purification systems more efficient and less expensive, and producing improved biofuels and pharmaceuticals more cost-effectively.
"We inserted our new polymers into lipid bilayers, and they transported protons just as well as natural proteins," said Ting Xu, a professor at UC Berkeley and faculty scientist for the Materials Sciences Division at Lawrence Berkeley National Laboratory.
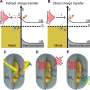
"The polymers are very difficult to image and study because of the limited contrast between their density and that of lipids. So we enhanced the contrast by selectively deuterating the lipids in the samples—meaning we replaced some of their hydrogen atoms with deuterium atoms—which neutrons are especially good at differentiating from hydrogen atoms. That allowed us to use neutron scattering at Oak Ridge to better 'see' the size and shape of the individual polymers," added Xu.
Working at ORNL's High Flux Isotope Reactor (HFIR), the researchers used the general purpose small-angle neutron scattering (GP-SANS) beamline to conduct their experiments.
"The GP-SANS instrument enabled the team led by researchers at UC Berkeley to determine the polymers were compact structures that were randomly dispersed within the membrane—as opposed to clumped together," said William T. Heller, the SANS/Spin Echo team lead at ORNL. "We chose the GP-SANS instrument because it is ideal for the polymer's size and its intense beam is excellent for studying samples that do not scatter strongly."
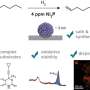
Xu and her collaborators said that the four monomers, the main components of the new polymer, can be grouped in different ways to produce functional protein-mimics. "What makes our new technique so promising is that it's scalable, and the knowledge to do this is readily available," said Xu. "Considering the vast number of monomers available and the recent advances in polymer chemistry, the possibilities of marrying the synthetic and biological fields are almost unlimited."
More information: Tao Jiang et al. Single-chain heteropolymers transport protons selectively and rapidly, Nature (2020). DOI: 10.1038/s41586-019-1881-0
Journal information: Nature
Provided by Oak Ridge National Laboratory