New software tool fosters quality control of genome-scale models of metabolism
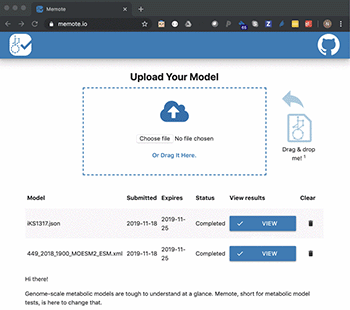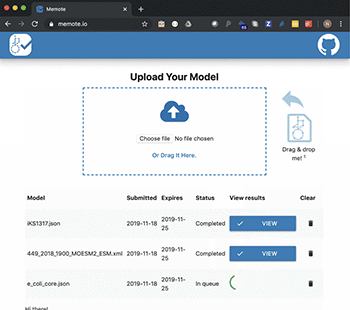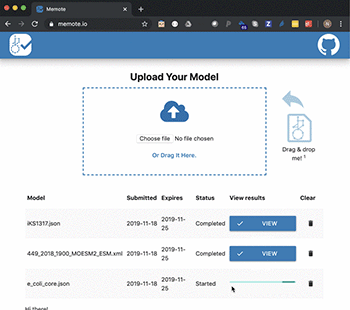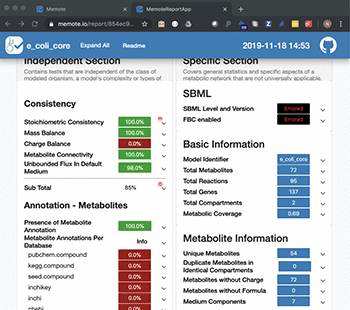
The application areas of genome-scale metabolic models are widespread, ranging from designing cell factories and investigating cancer metabolism, to analysing how microbes interact within our guts. Hence, the number of publications of manually and automatically generated models has been growing every year. This could be considered solely as a positive matter, but a lot of the data are very difficult for others to reproduce and reuse in different contexts.
Therefore, scientists from The Novo Nordisk Foundation Center for Biosustainability (DTU Biosustain) and a big group of researchers from all across the globe working within the field of biotechnology set out to address the problem.
In a new study published in Nature Biotechnology, they present the tool, MEMOTE, which is a community-maintained, standardised set of metabolic model tests. The tests cover a range of aspects from annotations to conceptual integrity and can be extended to include experimental datasets for automatic model validation.
Thus, the hope is that MEMOTE will allow both scientists and biotech companies to develop better performing models more efficiently. From a sustainability perspective, this will enable cell engineers to design and engineer better cell factories faster than currently possible since engineering the microbes for the production of biochemicals is a very time-consuming and expensive process at the moment.
Quality assurance in high demand
"If you work with model organisms such as yeast or the gut microbe Escherichia coli you are lucky because models for those organisms are quite predictive and have been refined over many years. But people in the community have been aware that a number of published models contain significant flaws," says Christian Lieven, who is a former postdoc at DTU Biosustain and now CSO and Co-founder of Unseen Bio.
Co-author Moritz Beber adds: "We conducted a quantitative assessment of thousands of published models. While the majority of them are in a reasonable state, MEMOTE was able to reveal specific problems in all of the models. This study underlines the utility of MEMOTE to assess and improve metabolic models".
Promotes openness and collaboration
Another issue that MEMOTE addresses by applying best practices from the field of Software Engineering, is the continuous improvement and versioning of models before and after publication. MEMOTE integrates with modern IT technologies such as the social coding platform GitHub and allows researchers to collaboratively improve models while making sure their quality never drops.
"Keeping a track record of model development is absolutely essential for both attributing credit but also for facilitating accountability. Furthermore, a lot of biological knowledge is uncovered from publications and documented in the process of reconstructing genome-scale metabolic models. Who knows, given the fact that reconstructions are knowledge-bases, maybe MEMOTE might even provide the means of publishing detailed reviews about organisms' metabolisms in the future," says Bernhard Palsson, co-author and CEO of DTU Biosustain.
Better cell factory engineering
MEMOTE enables a quick comparison of any two given models to assess which one is suited best for the selected host organism. A model is tested for a wide range of factors such as the production of biomass precursors, biomass consistency and growth rate. Ultimately, this should lead to a more rational approach to cell factory design.
"Today, we have so much data and knowledge about the biological pathways operating inside these industrial microorganisms. This enables us to use mathematical models to simulate the effects of genetic modifications and thus facilitate a more rational approach to the design of cell factories. I hope MEMOTE will facilitate the development of predictive models of many more organisms and thus also diversify the spectrum of available production hosts in cell factory engineering," says corresponding author and Associate Professor at DTU Bioengineering, Nikolaus Sonnenschein.
More information: Christian Lieven et al, MEMOTE for standardized genome-scale metabolic model testing, Nature Biotechnology (2020). DOI: 10.1038/s41587-020-0446-y
Journal information: Nature Biotechnology
Provided by Technical University of Denmark