Researchers reveal mechanism of trimming longer RNA transcripts to become micro RNA
A particular group of non-coding RNAs (ncRNAs) known as microRNAs (miRNAs) are ~22 nucleotide (nt) in length and play vital roles in diverse physiological processes. Aberrant expression of miRNAs has been linked to the development and progression of various human diseases including cancers. Certain miRNAs are even considered as biomarkers for diagnosis and targets for drug discovery.
The RNA transcripts destined for generating miRNAs have a characteristic stem-loop structure of ~65 nt in length, and the first step of becoming a functional miRNA is to have the extra RNA bits beyond the stem-loop region removed by a protein complex called Microprocessor in the cell nucleus. However, detailed mechanisms concerning the first cleavage reaction, including how pri-miRNA is recognized by Microprocessor, and how Microprocessor cuts at the correct sites on pri-miRNA, remain poorly understood.
Recently, a research team led by Dr. XU Ruiming from Institute of Biophysics of the Chinese of Academy of Sciences, and Dr. WANG Hongwei from Tsinghua University, determined a near atomic-resolution structure of Microprocessor bound to pri-miRNA. This study, published online in Molecular Cell, provided adequate answers to the mechanistic questions outlined above.
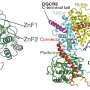
The Microprocessor complex is composed of Drosha, the catalytic subunit for RNA cleavage, and DGCR8, an RNA binding protein important for pri-miRNA binding. The researchers reconstituted the Microprocessor complex using purified proteins produced in insect cells, and pri-miRNA by in vitro transcription. The reconstituted RNA-protein complex or the protein complex alone were studied by single-particle cryo-electron microscopy to reconstruct the 3-D structure of the biomacromolecular specimen.
They found that Drosha plays an especially important role in pri-miRNA recognition and specification of the cleavage sites, in agreement with the results from previous studied. And they identified three key domains of Drosha that cooperatively recognize pri-miRNA, the PAZ domain, MB helices and the double-stranded RNA binding domain (dsRBD).
The PAZ and MB helices together recognize the single- to double-stranded junction region of pri-miRNA, and this interaction is critical for Drosha to bind pri-miRNA in the correct orientation and cut pri-miRNA at correct sites. Meanwhile, the dsRBD binds the upper stem region of pri-miRNA, stably positioning the RNA stem against to the catalytic center of the enzyme for cleavage reaction. The PAZ domain of Drosha binds both strands of RNA at the internal region, instead of the 3' termini bound by all other PAZ domains known to date.
Furthermore, they observed a large conformational change of the PAZ domain before and after RNA binding, suggesting an auto-regulated mechanism of RNA binding of Drosha, which may be beneficial for distinguishing specific targets from an abundance of various forms of RNA inside the cell nucleus.
This study solves a long-standing problem of how Drosha recognizes a pri-miRNA and binds it in a correct mode to allow precise cleavage of pri-miRNA. It provides fundamental understandings to the mechanisms of miRNA biogenesis.
In living organisms, genetic information stored in DNA, or genes, is first transcribed into messenger RNA by RNA polymerase II, and then the RNA transcripts direct the synthesis of proteins, which play the vast majority of the tasks in life processes. However, not all RNA transcripts are destined for making proteins, and those that do not code for protein synthesis are called ncRNAs.
More information: Wenxing Jin et al. Structural Basis for pri-miRNA Recognition by Drosha, Molecular Cell (2020). DOI: 10.1016/j.molcel.2020.02.024
Journal information: Molecular Cell
Provided by Chinese Academy of Sciences