SR-FACT microscopy reveals the landscape of the cellular organelle interactome
The emergence of superresolution (SR) fluorescence microscopy has rejuvenated the search for new cellular sub-structures and dynamic intermediates. However, limited by the broad emission spectrum of fluorophores and excessive phototoxicity, SR fluorescence microscopy can only be used to highlight a handful of biomolecules simultaneously and is incapable of providing a holistic map of the cellular environment and landscape.
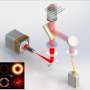
In a new paper published in Light Science & Applications, scientists from the State Key Laboratory for Mesoscopic Physics and Frontiers Science Center for Nano-optoelectronics, Peking University, China, State Key Laboratory of Membrane Biology, Beijing Key Laboratory of Cardiometabolic Molecular Medicine, Institute of Molecular Medicine, Peking University, China and co-workers developed SR fluorescence-assisted diffraction computational tomography (SR-FACT), which combines label-free three-dimensional optical diffraction tomography (ODT) with two-dimensional fluorescence Hessian structured illumination microscopy. The ODT module is capable of resolving mitochondria, lipid droplets, the nuclear membrane, chromosomes, the tubular endoplasmic reticulum and lysosomes. Using dual-mode correlated live cell imaging for a prolonged period of time, they observed novel subcellular structures named dark-vacuole bodies (DBs), the majority of which originate from densely populated perinuclear regions, and intensively interact with organelles such as the mitochondria and the nuclear membrane before ultimately collapsing into the plasma membrane. This work demonstrates the unique capabilities of SR-FACT, which suggests its wide applicability in cell biology in general.
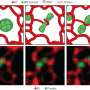
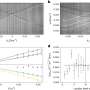
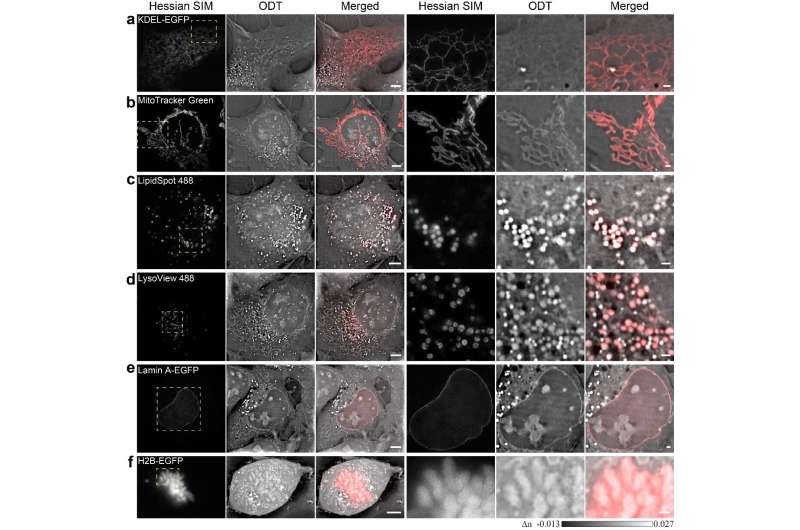
SR-FACT visualizes both the cellular landscape and the molecular identity of live cells. A new algorithm termed the vector iterative search algorithm (VISA) was developed to minimize 3-D imaging reconstruction errors under high speed kHz-rate tomographic scanning scheme. As a result, SR-FACT can simultaneously utilize a maximal imaging speed to capture dynamics in live cells and to maintain sufficient photon flux for maximal sensitivity. In the SR-FACT system, the ODT module achieved an ~200-nm lateral resolution at a volumetric imaging speed of 0.8 Hz (40×40×20 m3). Hessian 2-D-SIM, which allows SR imaging at a fraction of the photon dose used by conventional SIM, was used to guide the interpretation of structures observed by the ODT module. By performing dual-mode correlated imaging in COS-7 cells, they resolved six known organelles without labelling: the tubular endoplasmic reticulum (ER), mitochondria, late endosomes/lysosomes (LEs/LYs), LDs, the nuclear membrane and chromosomes. All these data highlight the unique advantage of SR-FACT in studying the organelle interactome. Moreover, they also observed vacuolated structures with neutral pH that contained mostly liquid in the lumen. Hour-long time-lapsed live cell SR imaging in combination with quantitative analysis reveals the unconventional trafficking routes and indispensable roles of vacuoles in organizing the organelle interactome, all of which suggests that they represent previously unappreciated organelles.
More information: Dashan Dong et al, Super-resolution fluorescence-assisted diffraction computational tomography reveals the three-dimensional landscape of the cellular organelle interactome, Light: Science & Applications (2020). DOI: 10.1038/s41377-020-0249-4
Journal information: Light: Science & Applications
Provided by Chinese Academy of Sciences