DNA nanostructures suit up for future missions
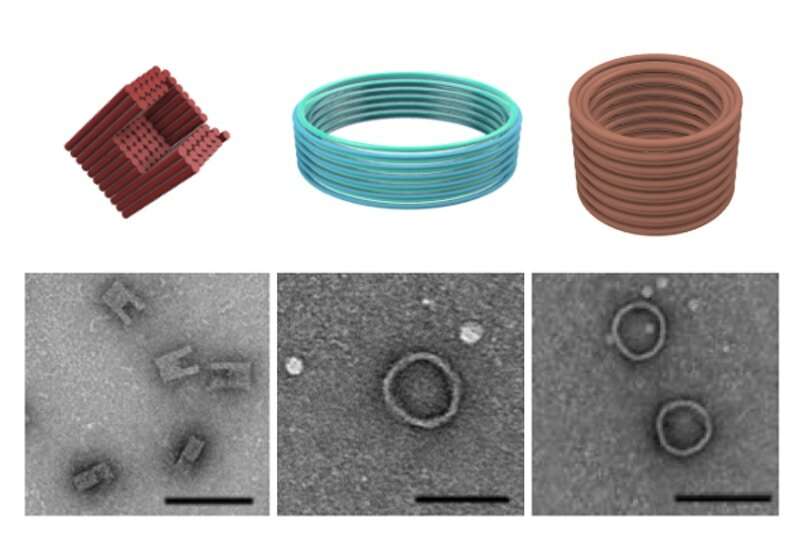

Beating cancer and a plethora of other diseases does not only depend on getting hold of the right drugs—it's also about getting them to the right places in the body, while keeping damage to healthy tissues at a minimum. DNA nanostructures such as self-assembling DNA origami that fold into defined shapes are promising vehicles for the delivery of drugs and diagnostics, and different vaccination strategies. This is because they can be flexibly functionalized with small molecule and protein drugs, and immunogenic antigens, as well as features that facilitate their delivery to specific target cells and tissues. In addition, they are biocompatible and biodegradable.
The great potential of DNA nanostructures for diverse biomedical applications, however, is still hampered by their limited stability in the body's tissues and blood. William Shih, Ph.D. and his team have now devised an easy, effective, and scalable chemical cross-linking approach that can provide DNA nanostructures with the stability they need as delivery vehicles for drugs and diagnostics. Their study is published in the Journal of the American Chemical Society (JACS).
"To help fulfill the extraordinary promise of DNA nanostructures, we developed a broadly applicable chemical approach to boost their resistance to degradation," said William Shih, a Core Faculty member at the Wyss Institute who is leading the project in the Institute's Molecular Robotics Initiative. "By modifying DNA nanostructures in two easy steps, we can dramatically enhance their survival, which removes a central bottle-neck in the translation of those nanodevices into solutions to a variety of medical problems."
DNA origami typically are assembled from a long strand of DNA that folds into a defined 3-D shape at the scale of nanometers with the help of multiple short staple strands connecting different parts of the structure in space. The DNA backbone is strongly negatively charged, which means that bringing different parts of the folding strands in the vicinity of each other can be problematic because they naturally repel each other, similar to the negative surfaces of two magnets. To overcome these repulsive forces, researchers commonly add an excess of positively charged ions like magnesium ions (Mg2+) to their folding reactions that blanket out DNA's negative charges.
Often, however, the added Mg2+ ions are not sufficiently maintained in 3-D DNA origami structures. Especially when introduced into the body, where normal Mg2+ concentrations are much lower, Mg2+ ions leak from the nanostructures and cause them to disintegrate. In addition, DNA nanostructures are under constant attack by DNA nuclease enzymes that are naturally produced in the body and degrade free DNA molecules, dramatically reducing their stability, which commonly is measured as their half-life. It logically follows that by disrupting the structural integrity of DNA nanostructures, these internal and external forces also curtail the release of drugs or availability of diagnostic reporter molecules.
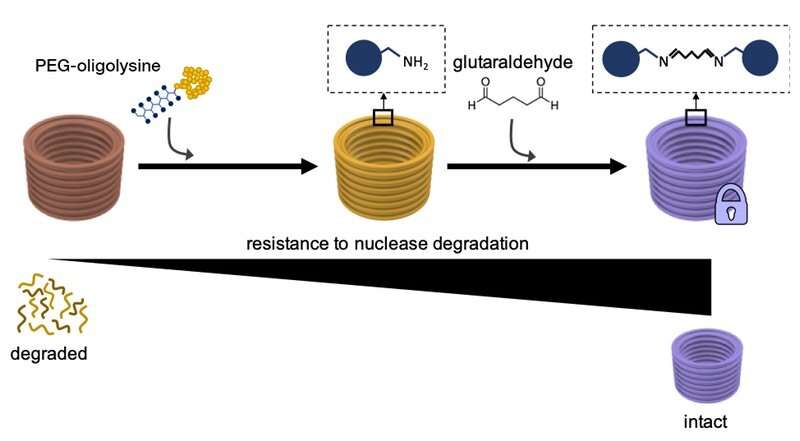
In a previous study, Shih and his team found a way to overcome the need for excessive Mg2+ concentrations to stabilize DNA nanostructures. By essentially coating the surfaces of the large strands of DNA with a comparatively tiny unobtrusive neutralizing agent that carries multiple positive charges and is known as PEG-oligolysine (or PEGylated oligolysine), they had invented a method that could be easily applied to stabilize DNA nanostructures in all shapes and forms in a scalable way. PEGylated oligolysine consists of short chains of the positively charged amino acid lysine—one of the building blocks of proteins—which are firmly attached to polyethylene glycol (PEG), a polymeric molecule able to bind multiple oligolysine chains. The researchers think that PEG functions as a "molecular brush" to prevent aggregation of the neutralized nanoparticles.
In contrast to individual Mg2+ ions that, in theory, each can neutralize only two of the many negative charges exposed by the DNA backbone with their two positive charges, PEGylated oligolysine can cover multiple negative DNA charges at once, and thus forms a stable "electrostatic net" covering the surface of DNA nanostructures. The stability of the coated nanostructures was even maintained in living cells and mice, demonstrating that the team was on the right track.
"Using this simple strategy, we could extend the survival of DNA nanostructures by about 400-fold with half-lives up to 36 hours in physiological environments that also contained normal nuclease levels," said first-author Frances Anastassacos, Ph.D., who at the time of the study was a graduate student working with Shih. "However, this still fell short of what is needed in some actual therapeutic applications. PEGylated oligolysine still tended to fall off the DNA to some degree."
In the JACS study, the team now solved this remaining part of the "stability problem." "We essentially tied additional knots into the electrostatic net using a chemical cross-linking reagent known as 'glutaraldehyde' that strongly bonds distinct PEGylated oligolysines on the surface of DNA nanostructures," said Anastassacos. "In the presence of DNA nuclease levels that far exceed those in the human body, their survival extended beyond 300 hours. The 400-fold increase in stability that we observed with PEGylated oligolysine alone was elevated by another 250-fold, which hits the mark for clinical applications."
Other attempts that succeeded in stabilizing DNA nanostructures with chemical modifications mostly relied on the re-design of existing structures so that specific sites within their 3-D structures would become accessible to the deployed chemistry. In contrast, "our approach can be performed on any DNA nanostructure, and its ease and cost-effectiveness could significantly strengthen the translation of many already existing and newly designed therapeutic and diagnostic structures," said Zhao Zhao, Ph.D., who is a co-first author on the study and former Postdoctoral Fellow on Shih's team.
In their study, Shih and his team aimed at maximizing the stability of DNA nanostructures. The researchers, however, think that by further varying the cross-linking step, the half-life of the structures could be fine-tuned to specific applications that require cargo molecules to become available at different times and over different time intervals. They already found that their protection method did not interfere with the binding of cargo molecules to the modified structures and the uptake of the structures by cultured human cells. Now, clinical translation can begin.
More information: Frances M. Anastassacos et al. Glutaraldehyde Cross-Linking of Oligolysines Coating DNA Origami Greatly Reduces Susceptibility to Nuclease Degradation, Journal of the American Chemical Society (2020). DOI: 10.1021/jacs.9b11698
Journal information: Journal of the American Chemical Society
Provided by Hansjörg Wyss Institute for Biologically Inspired Engineering