Direct microscopy imaging reveals details in DNA origami nanostructures
Scientists at the U.S. Department of Energy's Ames Laboratory are now able to see greater details of DNA origami nanostructures, which will lead to a greater understanding and control of their assembly for future applications.
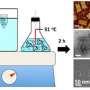
DNA origami is the folding of long "scaffold" strands of single-stranded circular DNA molecules held together using short "staple" strands to create various two- or three-dimensional shapes at the nanometer scale. DNA is widely used as building blocks for these structures because of specificity of the interactions between complementary base pairs. Because these nanostructures can be made identically and in relatively large quantities, they are very attractive to nanotechnology researchers, and are potentially useful in a number of applications.
Tanya Prozorov, an electron microscopy expert at Ames Laboratory, calls herself the "how do you know?" person. "One of the struggles of nanoscience is being able to clearly see the structures we are making, and understand these structures with greatest detail. How do we know? That is the problem I am trying to solve," she said.
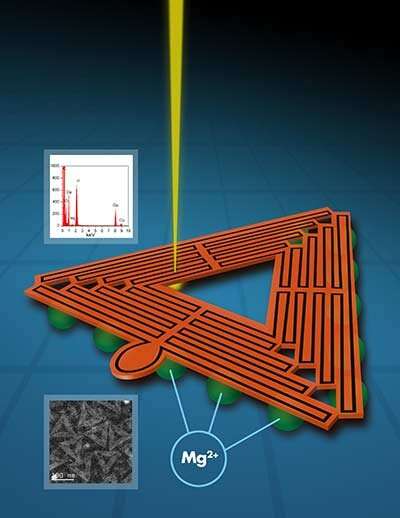
In this case, the challenge is to understand the three-dimensional size, shape and uniformity of triangle-shaped origami structures that will serve as a shape-specific foundation to create a layer of uniform gold nanoparticles, forming a plasmonic device, a metamaterial with specialized properties.
Due to their chemical makeup, the DNA nanostructures are difficult to image, with conventional electron microscopy sample preparation methods crushing or flattening the samples, or using a contrast stain that introduces other materials into the sample, possibly altering their surface chemistry and properties.
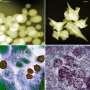
The Ames Lab research group was able to directly capture images of fine details in unstained DNA origami nanostructures using a particular type of scanning transmission electron microscopy called high- angle angular annular dark field STEM. Up until now, such level of detail of DNA origami structures was inaccessible without using chemical stains or resorting to atom force microscopy imaging. With the help of a collaborator from University of Pittsburgh, the team was also able to obtain a 2-D average image from a subset of lower-magnification images classified in a number of classes and calculated by using a specialized routine employed by the cryo-electron microscopy community.
"To make DNA-based plasmonic materials, we need to mount many metal nanoparticles on the DNA structures to achieve specific electromagnetic properties. The process begins at creating metal seeds at the surface of DNA origami, and so we need to have a good understanding of the surface—its structure, chemistry and topography, in order to understand the metallization process. Our method of imaging unstained, unaltered DNA origami structures allows us to see those details," said Prozorov. "We envision our method to be widely used as a straightforward method to image DNA nanostructures."
The research is further discussed in the paper, "Imaging of Unstained DNA Origami Triangles with Electron Microscopy," published in Small Methods.
More information: Alejandra Londono‐Calderon et al. Imaging of Unstained DNA Origami Triangles with Electron Microscopy, Small Methods (2019). DOI: 10.1002/smtd.201900393
Provided by Ames Laboratory