How to find molecular glues to effectively target diseases
Many of the currently available drugs are not specific enough to effectively cure complex diseases such as cancer, neurodegenerative diseases and diabetes. In addition, drug resistance reduces the effectiveness of existing therapies. To address these problems, biomedical engineer Eline Sijbesma designed small molecules that disarm specific disease proteins by gluing them to other proteins. These could lead to more stable and effective drugs and, among other things, might contribute to a new therapy for resistant breast cancer, for which currently no treatment exists. Sijbesma defends her Ph.D. thesis on December 2 at TU/e.
If you imagine the cell as the smallest factory of life, then think of proteins as the machines in these factories, doing all the work. Similar to machines in a production-line, proteins do not operate in isolation; they need each other to function. Physical interactions between proteins create essential signaling networks, enabling cells to quickly and adequately respond to external cues. In disease, often a single protein does not function well, or is too active. The activity of these disease-related proteins can be corrected by tightly binding them to small molecule drugs which can restore their normal function. However, this approach is not always successful and even if it is, diseased cells often find ways around the treatment.
Two is better than one
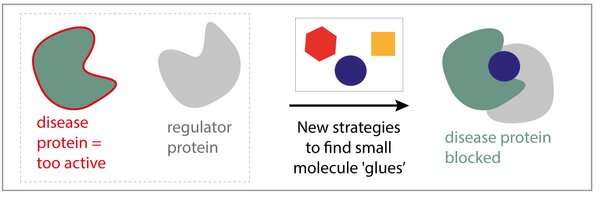
Eline Sijbesma, Ph.D. student in the research group of Chemical Biology led by Professor Luc Brunsveld: "Instead of focusing on a single protein and trying to find a specific drug for it, why don't we aim to make drugs that bind to a complex formed by two proteins?" Sijbesma's hypothesis is that if we can develop molecules that stabilize the interaction of a protein to another, we might be able to 'glue' a disease protein to a 'regulator protein' that prevents its bad activities. Additionally, these molecules are much more selective, since they only bind the protein complex, not the two individual proteins, resulting in less side effects.
Tight and strong
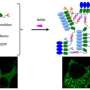
Together with the University of California San Francisco (UCSF) and the Novartis Institutes for Biomedical Research (NIBR), Sijbesma pioneered several drug discovery strategies making use of very small molecules (fragments) as starting points. Sijbesma: "We identified several fragments with desired properties and combined them together in a clever way, to form one new molecule with even better properties. We showed that the new molecule can indeed bind to two protein partners simultaneously, thereby making the protein complex up to 40-fold stronger."
Breast cancer
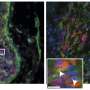
The development of these "molecular glues" has a great potential for certain types of resistant breast cancer. The latter often display an over-active receptor (Estrogen Receptor α), which is currently inefficiently targeted via drugs trying to block its activity directly. In a recent publication, Sijbesma proved that stabilizing the interaction of this receptor with a regulator protein might be the way to go. Sijbesma: "This receptor is known to be downregulated by a specific protein, the hub protein 14-3-3σ. Thus, we designed molecular glues that can tightly trap the receptor with this specific protein and inactivate it.
Beyond cancer
For Sijbesma, the central innovation of her research is the establishment of the underlying biological concept of a molecular glue for two proteins. This could lead to new avenues in drug development and the treatment of several diseases. Sijbesma: "This approach is not limited to applications in breast cancer, but it could be useful in the future for the development of novel therapies for diseases like neurodegeneration, inflammation, cystic fibrosis and diabetes."
More information: Eline Sijbesma et al. Site-Directed Fragment-Based Screening for the Discovery of Protein–Protein Interaction Stabilizers, Journal of the American Chemical Society (2019). DOI: 10.1021/jacs.8b11658
Title of PhD-thesis: 'Stabilization of protein-protein interactions as a drug discovery strategy; fragments as a muse'.
Journal information: Journal of the American Chemical Society
Provided by Eindhoven University of Technology