Estimates of commercial fish biomass from DNA traces in seawater
A new study published by scientists from the Faroese Marine Research Institute reveals that traces of DNA left behind in seawater can be used to predict the biomass of Atlantic cod. The findings show that the so-called environmental DNA approach can track the regional patterns of commercially important fish in the ocean. The paper, led by Dr. Ian Salter, was published on the 10th December in Nature Communications Biology.
All organisms continuously shed DNA into their environment, leaving behind a molecular footprint that can be used to identify the presence of different animal and plant species. Analysing these DNA traces represents a revolutionary approach for studying ecosystems that holds enormous promise. In particular, it has received a lot of attention in aquatic environments because of its potential to contribute towards fisheries management efforts.
The sustainable exploitation of commercial fisheries requires solid estimates of the quantity of fish stocks, and how they are changing over time and space. Traditionally, intensive bottom trawl surveys are used for this purpose and although they have been integral to decision makers, they have some notable drawbacks including cost, habitat destruction, species selectivity and restricted coverage. It is here that scientists believe environmental DNA can help out.
Previous studies in the last few years have shown analyzing DNA traces in seawater can describe the different species of fish caught in trawls. This is very useful for studying biodiversity and ecosystem dynamics. However, fisheries management relies on quantitative metrics such as Catch per Unit Effort, more specifically how many kg of fish are caught for each hour of trawling. The new study by Salter et al. describes the first example of how environmental DNA can be used in a quantitative way to provide these biomass metrics of commercially important fish in the ocean.
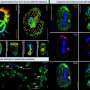
The authors mapped the distribution of Atlantic cod around the Faroe Islands from DNA traces in seawater samples, that were no larger than a fizzy drinks bottle, and compared it to standard trawl surveys. The two methods showed very good agreement both in the detection rate and quantities of Atlantic cod present in different regions. "It opens up the potential to track fish stock dynamics in the ocean independent of trawling vessels, comments lead author Dr. Ian Salter." This could be very useful in areas where trawling is difficult (shallow grounds or hard bottoms) or forbidden (coral reefs, marine protected areas). It also provides the opportunity to gather more information on fish stocks from a wider range of ships, and even autonomously using ocean robotics. "There is still some research required to understand precisely how many samples should be taken to get the most reliable estimates," says Dr. Salter, "but the approach has enormous potential."
More information: Salter, I., Joensen, M., Kristiansen, R. et al. Environmental DNA concentrations are correlated with regional biomass of Atlantic cod in oceanic waters. Nature Communications Biology 2, 461 (2019) DOI: 10.1038/s42003-019-0696-8
Journal information: Communications Biology
Provided by Faroese Marine Research Institute