Programmable swarmbots help make flexible biological tools
Biomedical engineers at Duke University have developed a new platform to create biologic drugs using specially engineered bacteria that burst and release useful proteins when they sense that their capsule is becoming too crowded.
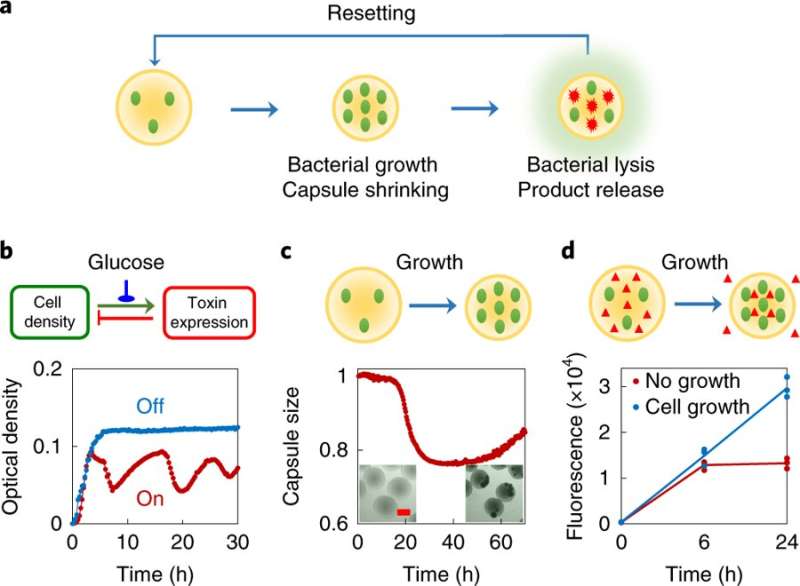
The platform relies on two main components: the engineered bacteria, called "swarmbots," that are programmed to sense the density of their peers within their container, and the biomaterial that confines the swarmbots, a porous capsule that can shrink in response to changes in the bacterial population. When it shrinks, the capsule squeezes out targeted proteins created by the captive bacteria.
This self-contained platform could make it easier for researchers to create, analyze and purify diverse biologics for use in small-scale biomanufacturing.
The research appeared online Sept. 16 in the journal Nature Chemical Biology.
Bacteria are commonly used to produce biologics, which are products like vaccines, gene therapies and proteins that are created or synthesized from biological sources. Currently, this process involves a series of sophisticated steps including cell culturing, protein isolation and protein purification, each of which requires delicate infrastructure to ensure efficiency and quality. For industrial operations, these steps are carried out on a large scale. While this helps produce large quantities of certain molecules, this setup is not flexible or financially viable when researchers need to produce small amounts of diverse biologics or work in resource-limited settings.
The new technology was developed by Lingchong You, a Professor of Biomedical Engineering at Duke University, and a former Duke postdoctoral researcher, Zhuojun Dai, now an Associate Professor at Shenzhen Institute of Advanced Technologies. In the new study, they show how their new platform uses communication between swarmbots and their capsule to achieve versatile production, analysis and purification of diverse proteins and protein complexes.
In an earlier proof of concept, You and his team engineered a non-pathogenic strain of E. coli bacteria to produce an antidote to antibiotics when the bacteria reached a certain density. These swarmbots were then confined to a capsule, which was bathed in antibiotics. If a bacterium left the capsule it was destroyed, but if it remained inside the container where the population density was high, it survived.
"Our first study essentially showed one-way communication, where the cells could sense the environment within the capsule but the environment didn't react to the cells," said You. "Now, we have two-way communication—the engineered swarmbots can still sense their density and their confinement, but we have introduced a material that can respond when the bacterial population inside it changes. It's like the two components are talking to each other, and collectively they give you very dynamic behavior."
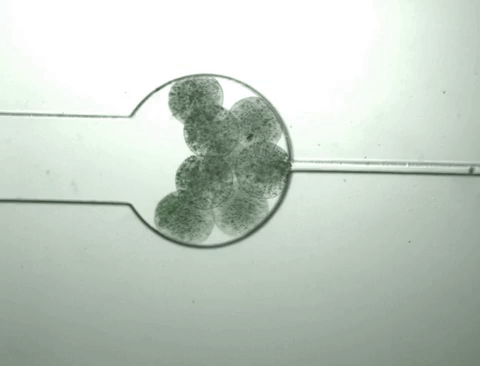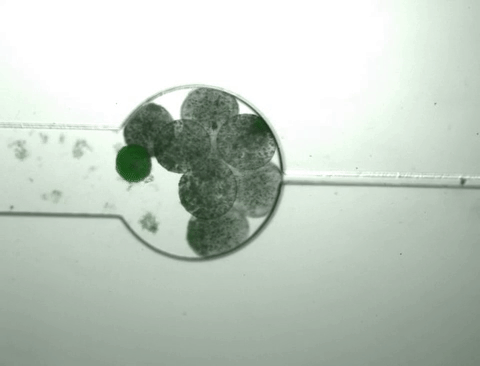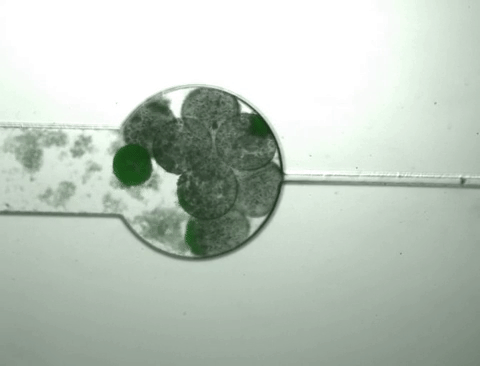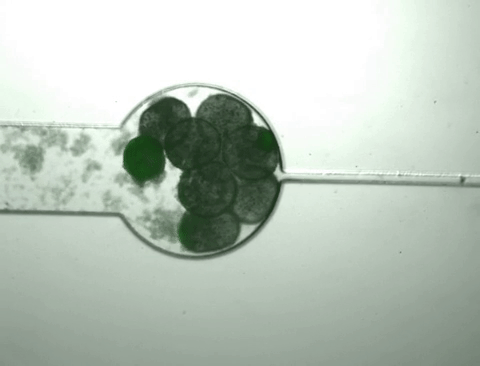
Once the population inside the capsule reaches a certain density, the bacteria start to 'pop,' releasing all of their cellular contents, including the protein product of interest. At the same time, this bacterial growth changes the chemical environment within the capsule, causing it to shrink. As it shrinks, it squeezes out the protein released from the bursting cells while the bacteria and cell debris are kept within the capsule.
Once the proteins are collected, researchers can add a nutrient replenishment to the dish as a cue for the capsules to enlarge. This resets the interior environment and allows the bacteria to begin growing again, restarting the process. According to You, this cycle can be repeated for up to a week.
To make the approach useful for biomanufacturing, the team added the capsules to a microfluidic chip, which included a chamber for them to detect and quantify which proteins were released. This could be replaced with a purification chamber to prepare the proteins for use in biologics.
"It's a very compact process. You don't need electricity, and you don't need a centrifuge to produce and isolate these proteins," said You. "It makes this a good platform for biomanufacturing. You have the ability to produce a certain type of medicine in a very compact format at a low cost, and it's easy to deliver. On top of that, this platform offers an easy way to produce multiple proteins simultaneously."
According to You, this ease of use has enabled the team to produce, quantify and purify more than 50 different proteins in collaboration with the lab of Ashutoshi Chilkoti, Alan L. Kaganov Professor and Chair of the Department of Biomedical Engineering at Duke. They have also explored how their platform can simplify the creation of protein complexes, which are structures made from multiple proteins.
On a proof-of-concept experiment to produce a fatty acid synthesis pathway from multiple enzymes, "we were able to use seven versions of our microbial swarmbots, each of which was programmed to produce a different enzyme," said You. "Usually, to produce a metabolic pathway you'd need to balance the supply chain, which could involve upregulating the expression of one enzyme and downregulating the expression of another. With our platform you don't need to do that, you just need to set the correct ratio of swarmbots."
"This technology is incredibly versatile," he said. "That's a capability we want to take advantage of."
More information: Zhuojun Dai et al. Versatile biomanufacturing through stimulus-responsive cell–material feedback, Nature Chemical Biology (2019). DOI: 10.1038/s41589-019-0357-8
Journal information: Nature Chemical Biology
Provided by Duke University