Entropy explains RNA diffusion rates in cells

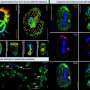
Recent studies have revealed that within cells of both yeast and bacteria, the rates of diffusion of RNA proteins—complex molecules that convey important information throughout the cell—are distributed in characteristic exponential patterns. As it turns out, these patterns display the highest possible degree of disorder, or 'entropy', of all possible diffusion processes within the cell.
In new research published in European Physical Journal B, Yuichi Itto at Aichi Institute of Technology in Japan explores this behaviour further by zooming in to study local fluctuations in the diffusion rates of RNA proteins. By associating these small-scale diffusion rates with time-varying values for entropy, he finds that the rates of change of entropy in certain time intervals are larger in areas with higher RNA diffusion rates.
Itto's work provides new insights into the complex biochemical processes that go on inside cells. This work could enable researchers to place more rigorous mathematical constraints on the ways in which they function. He also shows that the diffusive dynamics of RNA are analogous to thermodynamic behaviours in larger systems. His calculations imply that the differences in time-varying entropy in different parts of a cell are directly comparable to the time-varying differences in temperature resulting from the flow of heat throughout thermal systems. Itto derived these behaviours by using a series of mathematical equations. These relate RNA diffusion rates on small scales with their subsequent diffusion-varying rates of entropy.
Thanks to this approach, he has now successfully derived the characteristic exponential patterns of RNA diffusion rates, starting from basic mathematics. For the first time, his findings support previous observations that within yeast and bacteria cells, RNA diffusion represents the maximum possible distribution of entropy.
More information: Yuichi Itto, Time evolution of entropy associated with diffusivity fluctuations: diffusing diffusivity approach, The European Physical Journal B (2019). DOI: 10.1140/epjb/e2019-100054-9
Journal information: European Physical Journal B
Provided by Springer