July 24, 2019 report
Researchers develop phage-assisted continuous evolution of base editors system

A team of researchers from the Broad Institute, Harvard and Boston's Children's Hospital has developed a new way to improve the editing efficiency of base editors using a system called "phage assisted continuous evolution of base editors," or BE-PACE. In their paper published in the journal Nature Biotechnology, the group describes their new system and how well it worked.
The development of the CRISPR gene editing system has led to the possibility of editing genes to prevent inherited diseases. But problems with the system have persisted—most notably, instances of the wrong genes being edited. Because of this, scientists are seeking ways to improve the accuracy of such systems to make them safe enough for use in human patients. In this new effort, the team in Massachusetts has developed a system they call BE-PACE, which can be used to improve cytosine base editors (CBEs). The researchers used their system to evolve a CBE called evoAPOBEC1-BE4max. They report that testing showed it to be 26 times as efficient at editing cytosines (in the GC context) than prior systems, even as it maintained its efficiency in editing all other tested sequences. They further report that that testing of an evolved deaminase called evoFERNY showed it to be 29 percent smaller than APOBEC1.
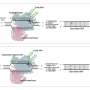
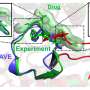
The researchers note that one of the factors that limit the efficiency of other CBEs is the context of native sequence preferences of APOBEC1, which leads to poor deaminating of GC motifs. To overcome this problem, the researchers used PACE systems because they are able to carry out multiple generations of selections, mutations and replications in a single day. Their goal was to create base editors with improved targeting abilities. They report that the BE-PACE circuit that they developed tested at almost ten-fold phage propagation in overnight host-cell cultures—and they demonstrated 1000 times the selectivity with base editor phages. The team also created another circuit to address sequence context limitations with APOBEC1. This led to the development of phage clones showing 28-fold improvements in activity during testing. To show that they were improvements for base editing, the team subcloned several evolved deaminase variants of a BE4max editor and inserted them into test cells using guide RNAs.
More information: Benjamin W. Thuronyi et al. Continuous evolution of base editors with expanded target compatibility and improved activity, Nature Biotechnology (2019). DOI: 10.1038/s41587-019-0193-0
Journal information: Nature Biotechnology
© 2019 Science X Network