Cancer treatment could become more effective thanks to new algorithms which can predict millions of gene interactions

While network algorithms are usually associated with finding friends on social media, researchers at the University of Sussex have shown how they could also be used improve the effectiveness of cancer treatment, by predicting the interactions between genes.
There are over 12 million newly diagnosed cases of cancer globally each year and this figure only continues to grow.
Existing treatments like chemotherapy involve non-selective agents that have limited effectiveness and strong side-effects. As a result, scientists believe there is a desperate need for improved treatments which are more personalised and more targeted towards cancerous cells.
There are a number of targeted cancer therapies already being developed that exploit a gene relationship called 'synthetic lethal interactions'. The trouble is, up until now, relatively few of these interactions have been identified.
Thanks to the use of artificial intelligence, researchers at the University of Sussex, working with a team from the Institute of Cancer Research in London, have successfully created an algorithm which can now predict where these interactions may occur.
Graeme Benstead-Hume, a doctoral student at the University of Sussex, said: "Synthetically lethal means that cells can cope if either one of its proteins does not work, but will die if neither of the proteins is functioning.
"These relationships are important because they can be used to identify where potential drug treatments could target just the cancer cells yet leave healthy cells unharmed, creating a more effective, gentler treatment.
"With breast cancer, we've already seen that these more personalised therapies can be achieved by finding synthetically lethal pairs of proteins. The only problem is that there are many millions of potential pairs and finding new ones is both difficult and time-consuming.
"Thankfully, our algorithm, Slant, can now address this."
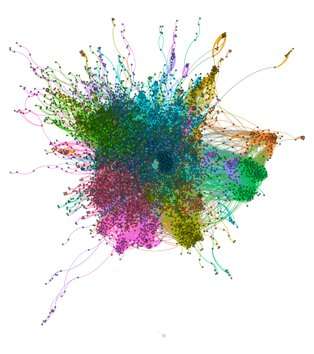
Spinglass showing complexity of human gene interactions
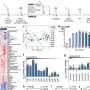
Slant uses data already available to identify patterns associated with being part of a synthetic lethal interaction. By searching across an expansive protein network for similar patterns, it's able to effectively predict new synthetically lethal pairs.
These predictions were validated by the researchers back in the laboratory and are now publicly available on a newly created database called Slorth, which allows clinicians and researchers to quickly search for a particular gene or drug, and identify whether a synthetic lethal interaction might occur.
This innovative computational approach has now been published in the journal PLOS Computational Biology.
Dr. Frances Pearl, corresponding author on the paper, said: "This work just shows how emerging technology and artificial intelligence can rapidly speed up the work that can lead to new treatment strategies for diseases like cancer.
"By predicting interactions between genes, we have sped up a process that would have been incredibly time consuming."
'Predicting synthetic lethal interactions using conserved patterns in protein interaction networks' by Graeme Benstead-Hume, Xiangrong Chen, Suzi Hopkins, Karen A. Lane, Jessica Downs and Frances M.G. Pearl was published online by PLOS Computational Biology on 17th April 2019.
More information: Graeme Benstead-Hume et al. Predicting synthetic lethal interactions using conserved patterns in protein interaction networks, PLOS Computational Biology (2019). DOI: 10.1371/journal.pcbi.1006888
Journal information: PLoS Computational Biology
Provided by University of Sussex