Methods for large protein crystal growth for neutron protein crystallography

The ability to grow large protein crystals is the single biggest bottleneck that limits the use of neutron protein crystallography in structural biology. Protein crystals need to have volumes in the region of at least 0.1mm3. Theoretically there is no particular reason why crystals of this size cannot be grown. If they can be, neutron protein crystallography can provide crucial information on the location of hydrogen atoms details relating to hydration hydrogen bonding and ligand interactions. This type of information is of direct relevance to academic and pharmacologically driven research in the life sciences.
The challenge is thus to achieve large crystal growth in a reproducible, time-saving, labour-saving way. It would be ideal if in the future, neutron crystallographers can, after suitable pre-characterisation work, submit their solutions to an automated or semi-automated platform that would allow the exploration of a large range of conditions in a highly systematic way and to allow users to monitor growth from their remote computers.
Ashley Jordan at the Institut Laue-Langevin (ILL) in Grenoble, France, has been investigating two new crystal growth methods: the development of a module that could allow larger scale automated approaches in the future (task 1), and a flow crystallization system (task 2).
Task 1: A module for automated large crystal growth exploration
This SINE2020 project has focused on the development of a temperature controllable multi-well module in which crystal growth can be optimized. The idea of designing this module was to scale up the approach so that multiple crystallization wells with individual (programmable) temperature control could be used to explore a wide range of growth conditions. A prototype module was made that consisted of a custom plate design containing 6 × 4 wells where the individual crystallisation experiments can occur. Each well can be adapted to different conditions, with each having independent temperature control. The wells are heated using Peltier heating elements with a temperature feedback system that allows each well to be heated and cooled over a temperature range of 4 degrees C to 60 degrees C, with an accuracy of 0.1 degree. The set-up was designed to allow allow crystal growth to be monitored and recorded photographically.
Ashley Jordan, Ryo Mizuta and John Allibon (who developed the software) have built and tested the prototype system. Crystallization tests have been carried out using trypsin and rubredoxin.
Post-SINE2020, the idea would be to make these modules "plug and play" so that a more extended 'robotic' approach could be used. Crystallogenesis runs could be removed by the user on completion and other runs installed using another module – the module would be the working unit of a larger array – with all being camera visualisable and providing time-lapse information to a user portal.
Task 2: Flow crystallization
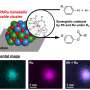
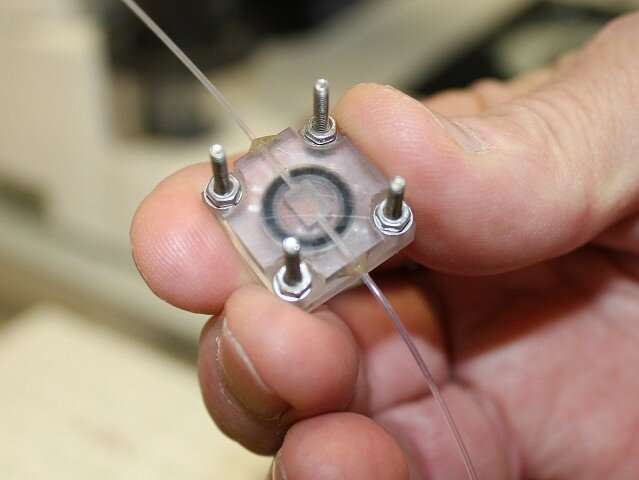
Another way of pursuing large crystal growth is the idea of a flow crystallization system. The idea is to maintain steady-state batch conditions around a crystal at all times during its growth, by providing a constant supply of fresh protein stock to the crystallization environment. This will maintain optimal solution conditions at all times and help minimize accumulation of impurities on crystal surfaces – such impurities may hinder crystal growth.
A Dolomite Mitos P-Pump was chosen to maintain the extremely low flow rate (between 70-1500 nl min-1) required to regulate the system. A suitable crystallization chamber that can connect to the pump was designed and made using a 3-D printer. This chamber creates a sealed environment and provides ready access to the crystals once they have grown.
Provided by CORDIS