Simplified method makes cell-free protein synthesis more flexible and accessible
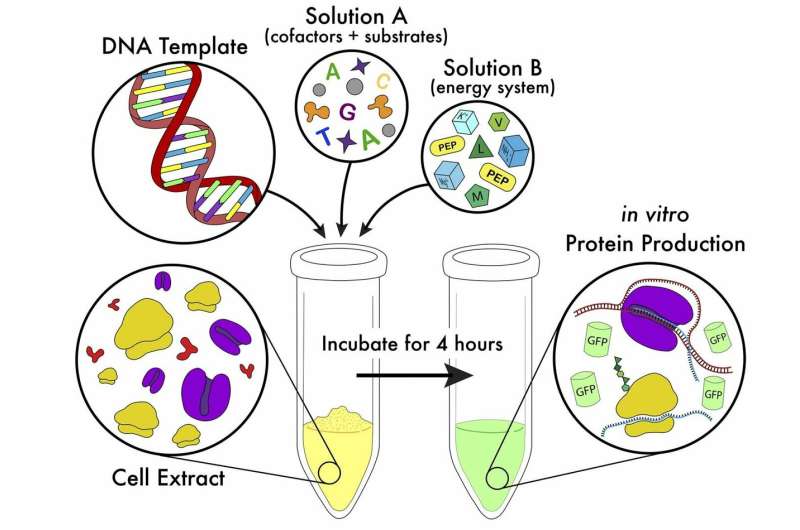
Researchers have radically simplified the method for cell-free protein synthesis (CFPS), a technique that could become fundamental to medical research.
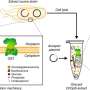
Synthesizing proteins is essential for multiple types of pharmaceutical and genetic research. For years, proteins could only be synthesized within live cells. CFPS provides the novel ability to biosynthesize proteins in a test tube in a matter of hours without the need for living cells. This process provides a new level of control over protein production, an incredible boon for researchers pursuing high-throughput testing, biosensor construction, metabolic engineering and more.
"This biotechnology harnesses the genetic code in a test tube, providing direct access to biological machinery that is traditionally locked inside the cell," said Javin Oza, a biochemistry professor at Cal Poly, San Luis Obispo. "This allows scientists and engineers to make vaccines and therapeutic proteins, and perform diagnostic tests on-demand in the lab or out in the field. In the classroom, CFPS allows students to learn about the genetic code in an inquiry-based manner."
While the acceptance of CFPS as a promising technology has grown substantially in the past two decades, it still suffers from some limitations due to the difficulty and expense of implementing the technique.
As reported in the Journal of Visualized Experimentation, a group of researchers at Cal Poly led by Oza and collaborator Katharine Watts have developed a method to make CFPS widely accessible. The main advantages of the new technique are speed, cost-effectiveness and a much less complex reaction setup compared to other CFPS systems. To further reduce the barriers for scientists to adopt this method, the publication includes a video guide for implementing the new procedure.
"This protocol simplifies and clarifies the methods for implementing cell-free protein synthesis by non-experts," Oza said. "Improved access to these methods will help democratize the platform and its broad set of applications."
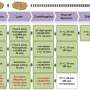
The new technique requires only basic laboratory training for new users to implement CFPS in their labs, from cell-growth and extract preparation to the in vitro protein synthesis reactions themselves. Researchers developed reagent premixes that greatly reduce the likelihood of error during setup and increase the probability of a successful reaction. These chemical and biological reagents are stable and can withstand multiple freeze-thaw cycles. In the paper, researchers also identify and report the variables that significantly impact the successful implementation of CFPS and the variables that new users should optimize for successful reactions.
The simplified procedure is functionally comparable to the more technical methods used previously and can be used to screen protein products more rapidly, greatly increasing the number of tests that can be run in a given time period. This capacity supports sophisticated research efforts, such as functional genomics and metabolic engineering.
"In our collaboration, we are employing the simplified CFPS method to synthesize and engineer complex mega-enzymes, which are traditionally difficult to express," said Watts, also a Cal Poly biochemistry professor. "The use of CFPS allows for much more rapid design-build-test cycles for engineering these mega-enzymes."
In addition to primary research, the new CFPS method can be utilized as an educational tool in high school and undergraduate classrooms. When packaged for educational use, it provides students the opportunity to engage with genetics in a hands-on learning environment.
More information: Levine, M. Z., Gregorio, N. E., Jewett, M. C., Watts, K. R., Oza, J. P. Escherichia coli-Based Cell-Free Protein Synthesis: Protocols for a robust, flexible, and accessible platform technology. J. Vis. Exp. (144), e58882, DOI: 10.3791/58882 (2019). www.jove.com/video/58882/esche … thesis-protocols-for
Provided by California Polytechnic State University