How bacteria mobility is like human locomotion

Do bacteria control their "walks" like we do? It might sound strange, but it's a fundamental question. Understanding bacteria motility would not only expand our understanding of their behavior, but would also contribute to the fight against certain aggressive pathogens. However, the question has gone unanswered because microbiologists have lacked the tools to visualize bacterial filaments directly.
Now, Lorenzo Talà, a Ph.D. student in the lab of Alexandre Persat at EPFL's Institutes of Bioengineering and Global Health has developed a microscopy method that can directly observe the structures many bacteria use to crawl. "Bacterial surfaces are decorated with protein filaments involved in motility, adhesion, signaling and pathogenicity, which ultimately rule how bacteria interact with their environments," says Talà. "However, they are so small that observing them in live cells is extremely complex. So we are left with little knowledge of their dynamic activities."
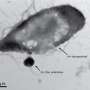
This is especially true for structures known as type IV pili: nanometer-wide filaments that extend and retract from the surface of many bacteria, helping them walk in a way known as "twitching motility." The term might not sound very serious, but it mechanically activates virulence in certain pathogens—meaning that it is a prime target for fighting them.
The scientists studied the bacterium Pseudomonas aeruginosa, an opportunistic pathogen that is commonly found in soil. It is a leading cause of hospital-acquired infections and of serious infections in cystic fibrosis, traumatic burns and immunocompromised patients, and is now ranked No. 1 in the World Health Organization's antibiotic resistance watch list.
But do single bacteria orchestrate type IV pili motion to power their motility? "In our studies of type IV pili and mechano-activation of virulence in Pseudomonas aeruginosa, one technical paradox has been a source of frustration: pili, fimbriae, flagella, and injection systems permanently extend outside single cells, so why can't we directly visualize them?"
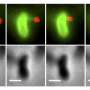
To overcome this, the scientists explored an emerging microscopy method pioneered by their collaborator Philipp Kukura at Oxford University. Using a technique called interferometric scattering microscopy (iSCAT), they were able to see these nanometers-wide filaments in live cells without any chemical labels, at high speed, and in three dimensions.
"iSCAT represents a major technological advance in microbiology," says Persat. "We recently described the visualization technique and received extensive positive feedback among scientists across a variety of disciplines simply because we could finally dynamically observe pili in live bacteria straight out of culture."
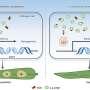
To understand the coordination of type IV pili movements, the scientists focused on precisely timing the succession of surface attachment, retraction, and cell body displacements using iSCAT. The approach revealed three key events that lead to successful and energetically efficient movement across surfaces.
First, contact of the pilus tip with the surface activates a molecular motor that initiates retraction. Second, this retraction enhances the attachment of the pilus to the surface, increasing the bacterium's displacement. Finally, a second, stronger molecular motor enforces the bacterium's displacement under high friction.
This sequence shows that pili act as sensors, and reveals a new mechanism by which bacteria interact with surfaces. It also reveals that bacteria use sensory mechanisms to coordinate the dynamic motion of their motility machineries, in a striking analogy to the way higher organisms, including humans, move their limbs to generate displacement.
"The human central nervous system processes mechanosensory signals to sequentially engage motor components, thus triggering muscle contraction and resulting in gait," explains Talà. "Our work shows that in the same manner, bacteria use a sense of touch to sequentially engage molecular motors, generating cycles of pili extension and retraction that result in a walk pattern."
The study is published in Nature Microbiology.
More information: Pseudomonas aeruginosa orchestrates twitching motility by sequential control of type IV pili movements, Nature Microbiology, DOI: 10.1038/s41564-019-0378-9 , www.nature.com/articles/s41564-019-0378-9
Journal information: Nature Microbiology
Provided by Ecole Polytechnique Federale de Lausanne