Mimicking nature for programmable and adaptive synthetic materials
Biological systems are truly representative of a complex macroscopic phenomenon, being acutely controlled by microscopic chemical reaction networks. As scientists seek to create a lifelike adaptive and responsive material, research has intensified into synthetic mimics of rudimentary biological processes.
One such biological process is the controlled growth of cytoskeletal proteins. Apart from acting as a nano-skeleton to maintain cell shape, assemblies of these proteins are responsible for distributing nutrients inside a cell. Thus, they control almost every important process inside the cell, from division to force distribution. It would be safe to presume that assembly control pathways of these proteins are a key contributor in a cell's adaptive and responsive behaviour. The key part of this process is an ATP-fuelled metabolic system that programmes the rate of growth and decay of these assemblies in a time-related manner. Close control is also maintained on the size of these assemblies as size directly relates to the functional efficacy of a system.
In a a study recently published in Nature Communications, scientists from the Jawaharlal Nehru Centre for Advanced Science and Research (JNCASR) and the Institute for Stem Cell Biology and Regenerative Medicine (inStem) created a minimalistic synthetic mimic of cytoskeletal networks with structural and temporal programming. The work focuses on reaction-driven controlled growth of a two-component monomeric molecular system.
"This synthetic monomeric system is elegantly designed so as to give rise to a primary reaction that would convert an inactive (non-assembling) monomeric system into an active one (which triggers assembly) upon addition of alkyl amines (fuel)," said Dr. Subi George, associate professor at JNCASR.
They show that this reaction can be used to precisely control the one-dimensional growth (nanofibers) of resulting assemblies through controlled availability of fuel. The growth of these nanofibrous structures is driven by very weak intermolecular interactions (supramolecular polymerization) such as hydrophobic reactions similar to lipid bilayers in membranes and aromatic interactions similar to stacking of nucleobases in DNA, and hence are highly dynamic and have self-repairing features like many biological assemblies.
"While biological systems elegantly modulate self-assembly with a great precision, imparting transient and living polymerization properties in chemical amphiphiles has been a daunting task thus far. Design of an in-situ forming amphiphile enabled us to study the dynamic assembled structures at will," said Dr. Praveen Kumar Vemula.
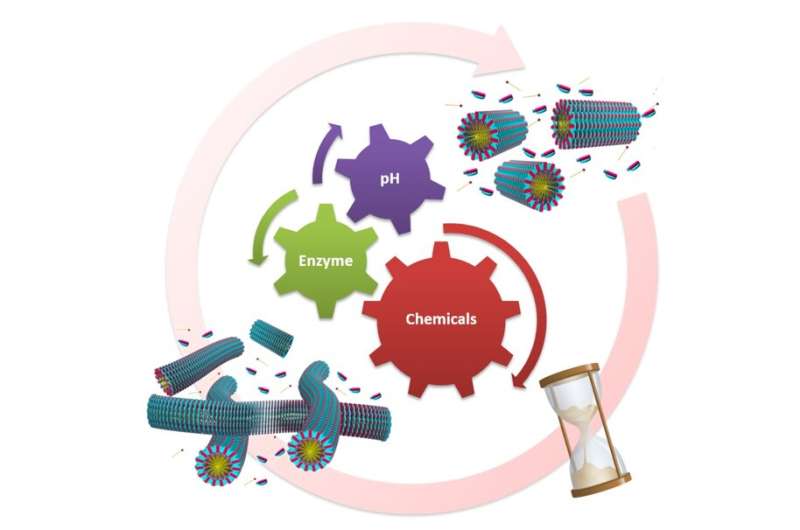
Through detailed spectroscopic and microscopic analyses, they established this growth to be "living" in nature, resulting in assemblies with very narrow size distribution (monodispersed). The system was further developed by coupling the growth with uniquely chosen chemical scenarios such that control over the growth and disassembly kinetics were established. As a result, a time-programmed transient network of fibrous assemblies was realized. In both cases, the manipulation of key temporal characteristics was from the range of a few seconds to thousands of seconds. This study thus represents a key step in the development of adaptive, lifelike, supramolecular materials.
"We have, for the first time, demonstrated that every temporal characteristic of supramolecular polymerization can be chemically controlled and further coupled to other participating reactions similar to a biological system," said Ankit Jain, lead author of the paper.
"Controlling the self-assembled nano-architectures using stimuli such as enzymes and pH has been fascinating," says Ashish Dhayani, author of the paper.
"This work is a significant advancement to design bio-mimetic active systems operating under out of equilibrium conditions, with spatio-temporal programming compared to the majority of the synthetic passive systems reported so far, which work under thermodynamic equilibrium with only spatial complexity," said Shikha Dhiman, co-author of the paper.
The next challenge is to build synthetic lifelike systems that can think, learn and adapt as living beings do. This study is one such initial step, but there is still much investigation required for fully emulating natural processes. The team hopes to apply this principle and use these dynamic self-assembly nano-architectures in biological systems.
More information: Ankit Jain et al, Chemical fuel-driven living and transient supramolecular polymerization, Nature Communications (2019). DOI: 10.1038/s41467-019-08308-9
Journal information: Nature Communications
Provided by National Centre for Biological Sciences