Increased speed of engineered human cell response
Researchers from the National Institute of Chemistry in Slovenia have developed a novel approach to regulate human cell response, endowing cells to respond to extracellular stimuli in minutes instead of hours. The authors believe that their system could be useful in various medical applications.
Cells function on the basis of a large number of processes working in conjunction to detect and respond to diverse stimuli. The control of cellular response is important in cellular therapy, as it could improve its efficiency and safety. Synthetic biology treats cells as machines or computers that we can edit and reprogram to our needs. While advances in genetic engineering have allowed precise sensing and programmed responses to biological and chemical inputs, the speed of information processing within cells until recently remained challenging. Researchers from the National Institute of Chemistry in Slovenia have now published an article in which they report that they were able to detect the cells responding to programmed inputs within minutes instead of hours.
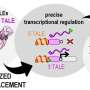
Ph.D. student Tina Fink, lead author on the study published in Nature Chemical Biology says, "Usually, biological circuits are based on gene expression regulation. This means that any input or change in environment first results in transcription of genes from DNA to RNA, and later RNA is translated into proteins. This process can take several hours. On the other hand, our system is based on processing of proteins already present in the cells, allowing us to circumvent the slow processes of transcription and translation."
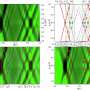
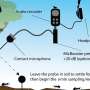
Two important processes that control protein activity in nature are protein cleavage and protein assembly into higher-order complexes. For example, researchers often use a luciferase protein split into two parts that can interact with each other, resulting in a functional complex that catalyzes a reaction that produces light. Dr. Roman Jerala, project leader and head of the Department of Synthetic Biology and Immunology, says, "Our team used proteases (proteins that cleave other proteins at specific positions) and combined them with coiled coils, which act as a sort of molecular zipper and allow for controlled protein coupling. The proteases were modified so that they become active only in the presence of a defined chemical signal or light. The action of the proteases induces the cleaving, and in turn, rearrangement of target proteins and correct processing of the information can be detected by light measurements. In this way, we can turn the activity of proteins on and off."
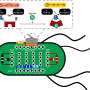
Although all the experiments measured light emission as a functional output, the final goal of the research is to control secretion of medically important proteins, such as insulin or proteins involved in the defense against pathogens or in the coagulation cascade. Dr. Jan Lonzarić, who shares first authorship with Tina Fink and is currently working as a postdoctoral fellow at Massachusetts Institute of Technology, said that after the speed of the reaction, the next most important property of this system is the ability to design parallel or sequential reactions, like logic gates and cascades. "The proteases we used to control protein assembly can also be controlled in the same manner, just using another protease. We successfully showed a three-layer cascade and logic gates that allow a programmed response to a combination of various inputs."
The project, which led to the publication in the journal Nature Chemical Biology, was first conceived by the Slovenian iGEM team in 2016. The group of students participating at the International Genetically Engineered Machines competition under the mentorship of Jerala and his group designed an initial concept for the system. "The project landed us first place in the Foundational Advance Track at the 2016 iGEM competition, but our goal was to continue developing this initial concept and, eventually, publish it, for which a lot of additional research was needed." said Arne Praznik, who joined the Department for Synthetic Biology and Immunology after iGEM to continue working on the project as a Ph.D. student.
Only a few weeks before the Slovenian team's research was published, another group from Caltech published a paper in the journal Science, which also approached the activation of human cells with the help of proteases. Lonzarić believes that simultaneous publications from different universities and institutes prove that "we have tackled an important problem and found a robust solution," while Jerala adds, "The differences between the two designs make our system faster and easier to employ in different cell types."
More information: Tina Fink et al. Design of fast proteolysis-based signaling and logic circuits in mammalian cells, Nature Chemical Biology (2018). DOI: 10.1038/s41589-018-0181-6
Journal information: Nature Chemical Biology , Science
Provided by National Institute of Chemistry, Slovenia