Researchers identify key players in mysterious process of protein quality control

Proteins are the workhorses of our cells, carrying out essential tasks to keep our cells – and our bodies – functioning properly. But proteins can only do their jobs if they fold into the right shape.
When a protein misfolds, the cell can try to salvage the situation by re-folding the protein or destroying it, but how cells make that decision has been a mystery. In a recent study published in Nature, Judith Frydman and her team identified the key molecular players in this decision. She sees this basic knowledge as the first step toward treating many human diseases, including neurodegenerative disorders like Alzheimer's and Parkinson's diseases and cancers that sometimes result when a cell fails to eliminate misfolded proteins.
"We need to understand how cells make this decision at the molecular level before we can develop cures," she said. "But first we had to define the components and pathways."
To re-fold or not to re-fold?
The trick is finding the right balance between eliminating bad proteins and salvaging what's fixable.
"The cell needs to find a quality control sweet spot where it degrades the misfolded proteins to keep them from accumulating and being toxic but isn't too zealous in degrading everything –including proteins that can still function," Frydman said.
After about a decade of work, Frydman and her team found how the cell determines where that sweet spot lies. The responsibility resides with two groups of proteins – called ubiquitin ligases and molecular chaperones – that work together to decide what to do about misfolded proteins.
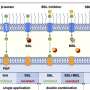
The ubiquitin ligases stick a variety of branched ubiquitin chains to specific locations on the misfolded proteins. Depending on the location and type of tags, the cell decides whether to destroy or re-fold.
Both groups of proteins are known to play other roles in the cell, but this is the first evidence that they have a dual role in targeting misfolded proteins. "Molecular chaperones, much like human chaperones, help newly made or otherwise immature proteins attain their mature, functional state," Frydman said. "I was surprised that chaperones also form part of the decision to degrade a misfolded protein."
Compartmentalizing the decision
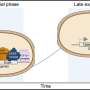
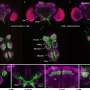
While studying how cells determine which misfolded proteins to destroy, the team noticed something strange: the selection processes differed in the cytoplasm, the main compartment of the cell, and the nucleus, where the cell's DNA resides.
"We think this reflects the different requirements for how stringent protein quality control has to be," Frydman said. In the cytoplasm – where newly created proteins are in the process of folding – cells would try not to degrade proteins prematurely. Cells might lower the bar for destroying misfolded proteins in the nucleus, to avoid them interfering with important genetic information.
Frydman and her team are still curious about how cells decide what to do with misfolded proteins in different parts of the cell, but their new findings have laid the foundation to answer questions like these.
"This basic knowledge establishes a new principle that will allow us to think about this important question of protein quality control in healthy cells but also during disease, both fundamentally and from the point of view of therapeutics," she said.
More information: Rahul S. Samant et al. Distinct proteostasis circuits cooperate in nuclear and cytoplasmic protein quality control, Nature (2018). DOI: 10.1038/s41586-018-0678-x
Journal information: Nature
Provided by Stanford University