Unveiling the mechanism protecting replicated DNA from degradation

Researchers from Tokyo Metropolitan University and the FIRC Institute of Molecular Oncology (IFOM) in Italy have succeeded in depleting AND-1, a key protein for DNA replication, by using a recently developed conditional protein degradation system. They have gained unprecedented access to the underlying mechanism and determined that during DNA replication and cell proliferation in vertebrate cells, AND-1 has two different functions mediated by different domains of AND-1.
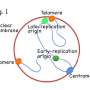
In order for living organisms to function, it is vital that all cells share the same DNA blueprint. This is made possible by the process of DNA replication, where the DNA is accurately copied and distributed before the cell multiplies. Replication underpins all biological inheritance, and is supported by a whole range of biochemical pathways designed to ensure that it occurs without error and at the right speed. Failure to do so can have catastrophic consequences, including cancer. Understanding the specific mechanisms behind this highly complex procedure is of the utmost importance.
The AND-1/Ctf4 protein is a key player in DNA replication, and is found in a vast range of living organisms. Ctf4/AND-1 is essential in some organisms, but whether it is an essential gene for cell proliferation in vertebrates has not been shown experimentally. Moreover, how loss of AND-1 affects cell proliferation is not known.
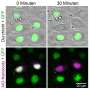
In order to address this question, a team led by Dr. Dana Branzei from IFOM and Prof. Kouji Hirota from Tokyo Metropolitan University combined the DT40 cell, a type of avian cell that is particularly suited to genetic engineering, and the auxin-inducible degron (AID) system, a means of selective depletion of a target protein. They established the and-1-aid cell line, in which a modified version of the AND-1 protein is degraded in a few hours after adding auxin, a type of plant hormone. This cell line expressed the acute consequences of AND-1 loss, giving unprecedented insight into the role the protein plays.
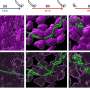
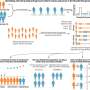
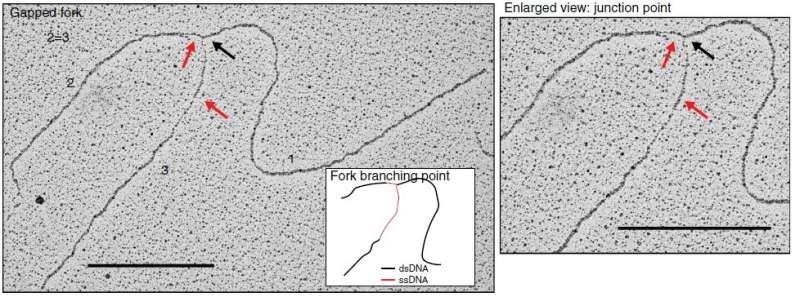
When done correctly, DNA replication should result in the formation of new double-stranded DNA helices. The authors used transmission electron microscopy (TEM) to visualize DNA replication intermediates and observed newly synthesized DNA with abnormally long single-stranded DNA at the fork branching point in the absence of AND-1. They hypothesized that this was due to a DNA cleaving enzyme, a nuclease, disrupting the process of strand disassembly. On further addition of a compound that suppresses the action of a particular nuclease, MRE11, they were able to revert the abnormal replication fork phenotype and recover cell division, explicitly demonstrating the key role played by AND-1 in preventing nascent DNA cleavage by the nuclease during replication. Further analysis revealed that a specific part of the protein called WD40 repeats was responsible for preventing the accumulation of damage to the strand.
The study highlights the successful combination of cutting-edge techniques to realize conditional inactivation of specific proteins; the new cells were developed over a single month. This suggests the exciting prospect of applying the method to study other genes and processes that are otherwise difficult to target, leading to new insights into how cells work.
More information: Takuya Abe et al, AND-1 fork protection function prevents fork resection and is essential for proliferation, Nature Communications (2018). DOI: 10.1038/s41467-018-05586-7
Journal information: Nature Communications
Provided by Tokyo Metropolitan University