New protein sequencing method could transform biological research
A team of researchers at The University of Texas at Austin has demonstrated a new way to sequence proteins that is much more sensitive than existing technology, identifying individual protein molecules rather than requiring millions of molecules at a time. The advance could have a major impact in biomedical research, making it easier to reveal new biomarkers for the diagnosis of cancer and other diseases, as well as enhance our understanding of how healthy cells function.
The team published the results of their proof-of-concept study today in the journal Nature Biotechnology.
"We have created, essentially, a DNA-sequencing-like technology to study proteins," said Edward Marcotte, professor of molecular biosciences and co-inventor of the new technology.
Work on this project began more than six years ago when Marcotte and colleagues first envisioned adapting the methods of next generation gene sequencing to protein sequencing. Next generation gene sequencing is a set of techniques that have made sequencing the entire genome of any living organism fast, accurate and affordable, accelerating biological research—and for the rest of us, enabling at-home genetic testing for ancestry and disease.
In the same way that these earlier advances provided quick and comprehensive information about thousands of genes that influence human health, the new technology provides for rapid and comprehensive information about tens of thousands of proteins that play a role in healthy functioning or in diseases. In many disorders—such as cancer, Alzheimer's, heart failure and diabetes—cells produce proteins and other substances that act as unique biomarkers, akin to fingerprints. Better detection of these biomarkers would help researchers understand the causes of disorders or provide earlier, more accurate diagnoses for patients.
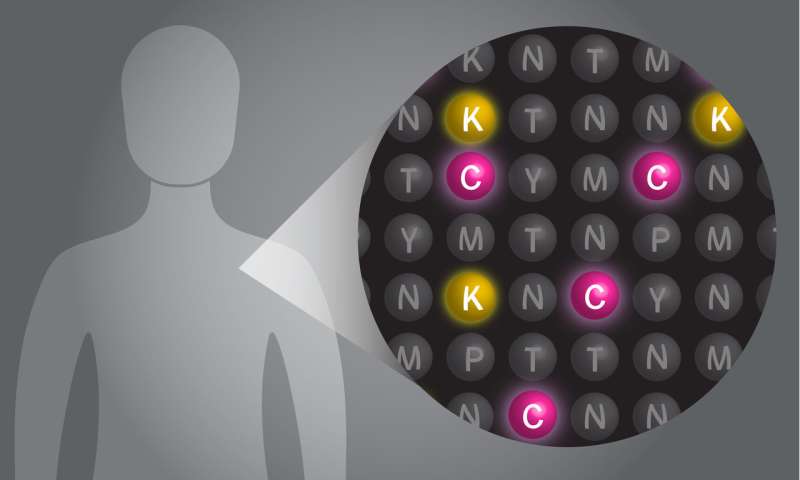

The current laboratory standard for sequencing proteins, using a tool called mass spectrometry, is not sensitive for many applications—it can detect a protein only if there are about a million copies of it. It also has a "low throughput," meaning it can detect only a few thousand distinct protein types in a single sample.
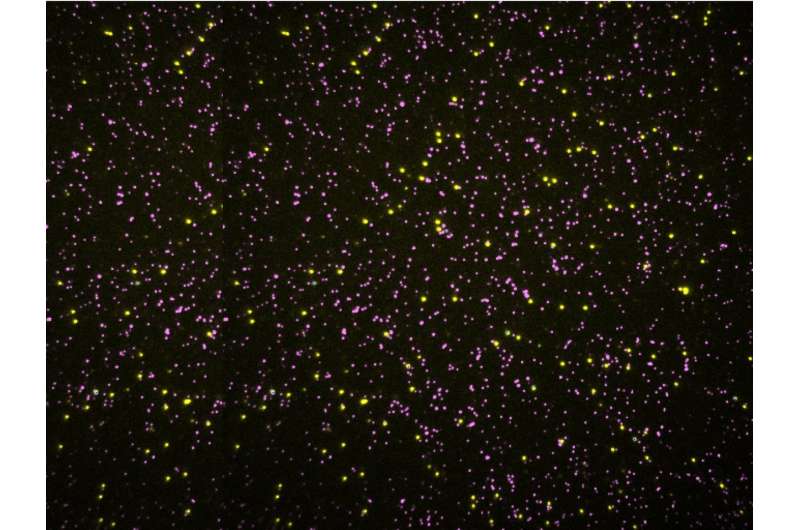
With this new method, called single molecule fluorosequencing, researchers can now sequence millions of individual protein molecules simultaneously in a single sample. Marcotte believes with future refinements the number of molecules that could be detected in a sample could reach into the billions. With higher throughput and much greater sensitivity than existing technology, the tool should allow for greater detection of biomarkers of disease and would also make it possible to study things such as cancer in a whole new way. For example, researchers could look, cell-by-cell, to understand how a tumor evolves from a small mass of identical cells to a soup of genetically divergent cells, each with its own strengths and weaknesses. Such insights could inspire novel ways to attack cancer.
More information: Highly parallel single-molecule identification of proteins in zeptomole-scale mixtures, Nature Biotechnology (2018). DOI: 10.1038/nbt.4278 , www.nature.com/articles/nbt.4278
Journal information: Nature Biotechnology
Provided by University of Texas at Austin




















