Bioinformaticians examine new genes the moment they are born
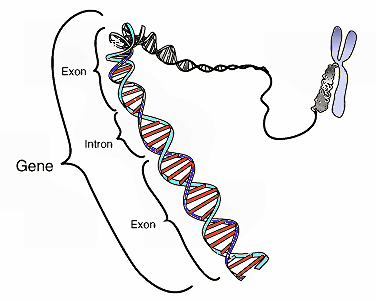
Accumulating evidence suggests that new genes can arise spontaneously from previously non-coding DNA instead of through the gradual mutation of established genes. Bioinformaticians at the University of Münster (Germany) are now, for the first time, studying the earliest stages in the emergence of these "genes out of thin air," also known as de novo genes.
Organisms evolve gradually due to the accumulation of many small genetic changes known as point mutations. Over millions of years, these mutations occur in the duplicated copies of established genes, occasionally contributing useful properties of their own. For decades, it was considered inconceivable that completely novel genes could emerge spontaneously. Only very recently were there serious indications that novel protein coding genes might indeed be formed de novo from so-called non-coding DNA, i.e. in parts of the genome that do not produce proteins. Now, for the first time, a new study has examined the earliest stages in the emergence of these de novo genes. The study, which has been published in Nature Ecology and Evolution, was carried out by a team of bioinformaticians led by Prof. Erich Bornberg-Bauer from the Institute of Evolution and Biodiversity at the University of Münster, Germany.
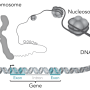
Using computer analyses, the team compared several properties of de novo genes in mice with those in four other types of mammals: rats, kangaroo rats, humans and opossums. Based on this comparison, the researchers were able to shed light on 160 million years of evolution in mammals. They took a close look at DNA transcripts (sequences present in the cell as RNA templates) that contain the open reading frames (ORFs) necessary for the encoding of proteins.
"Our study shows that new ORFs—in other words, the candidates for assembly instructions for new proteins—constantly emerge 'out of nowhere' in non-coding DNA regions," says bioinformatician Erich Bornberg-Bauer. "But, just like their transcripts, the vast majority disappear again very quickly during the evolutionary process." Although only very few of these candidates actually become fully functioning genes—i.e. genes containing the assembly instructions for functioning proteins—some of the candidates are retained at random for longer periods of time, simply because of the enormous number of new transcripts being continuously produced. "These transcripts can then be found in several lineages," says Bornberg-Bauer. "Probably, they can augment the repertoire of existing proteins over longer periods of time and become adapted to the interaction with such established proteins."
This means that a de novo protein can occasionally acquire a function in the organism. "This also provides us with an explanation of how fundamentally new properties can emerge in an organism," says Bornberg-Bauer, "because this cannot be explained just by point mutations in the genetic structure."
More information: Jonathan F. Schmitz et al, Incipient de novo genes can evolve from frozen accidents that escaped rapid transcript turnover, Nature Ecology & Evolution (2018). DOI: 10.1038/s41559-018-0639-7
Journal information: Nature Ecology & Evolution
Provided by University of Münster