Technique to fine-tune two-component biological sensors
Rice University scientists who say biological sensors aren't sensitive enough are doing something about it.
The lab of synthetic biologist Jeffrey Tabor has introduced a new technique to dial up or down the sensitivity of two-component systems – a class of proteins that bacteria use to sense a wide variety of stimuli.
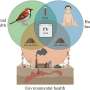
The technique could enable the engineering of tailor-made biosensors for diagnostic gut bacteria, detection of environmental pollutants or automated control of nutrient levels in soil.
Two-component sensors, the focus of a new paper describing the work in Nature Communications, are a large family of genetically encoded sensors that bacteria use to sense a specific input and turn on a specific gene in response to changes in their environment.
While these sensors have been known for three decades, they weren't sensitive enough for Tabor. He and Rice alumnus and lead author Brian Landry set out to decrease the amount of input needed to turn them on.
By combining computer modeling and experiments, they achieved just that. In the paper, they describe their discovery that phosphatase activity, which is essential to regulation and signaling in cells, can have a dramatic effect on the detection thresholds of two-component systems.
Previous research had shown that mutations to the first component, a signaling sensor protein known as a histidine kinase, can be used to control the extent of phosphorylation of the second component, a response regulator protein. But no one had suggested that phosphatase mutations might be used to alter the sensitivity of these pathways for their inputs, Tabor said.
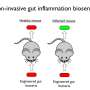
One day, faced with a nitrate sensor that was not turning on as expected in the gut of a sick mouse, Landry hypothesized that phosphatase mutations might dramatically increase the pathway's sensitivity.
He first validated the hypothesis using a mathematical model and then conducted experiments that showed the mutations increased the efficiency of a nitrate sensor by a factor of 100. Tabor's lab went on to validate the approach in two-component systems that sense a wide variety of inputs, even in very different species of bacteria.
Landry worked with another Rice bioengineering student, co-author Lucas Hartsough, to identify a "hot spot" amino acid present in 64 percent of all sensor histidine kinases that can be mutated to tune sensitivity. They validated the approach in two minimally characterized pathways, which suggested it can be widely applied, Tabor said.
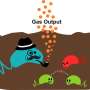
Landry also used the technology to engineer a soil bacterium, Bacillus subtilis, to sense and respond to a wide range of nitrate (fertilizer) concentrations in soil. Tabor and Landry suggest this system could be coupled to pathways under development by a colleague at MIT, synthetic biologist Chris Voigt, and his lab to engineer soil bacteria that maintain optimal nitrogen levels in soil without the need for fertilizer.
Two-component systems can be triggered by many kinds of inputs, including gases, heme molecules in blood, sugars, gut polysaccharides, human or plant hormones or even light.
"That's why we're so excited about this," Tabor said. "These sensors all operate the same way. They all have the phosphatase and the kinase. We've identified over 25,000 of them in bacterial genomes and we think our strategy will work on most of them."
He said his lab's technique relies on the signaling pathway by which natural sensor molecules tell cells to stop, start or modify the production of proteins. Tuning natural sensors for synthetic applications has been difficult until now, he said.
"One of the great limitations is that when you take a biosensor from nature, it may sense the chemical you're interested in, but not at the right concentration. This is because it has evolved to allow a bacterium to survive better in a natural environment rather than to satisfy the engineering needs of a synthetic biologist," he said.
"We call this level the detection threshold of the sensor, and there haven't been many good technologies to tune that threshold to match our engineering needs," Tabor said. "That's been a major limitation in biosensor design. But now, we've found one general trick that allows us to tune the detection thresholds of this family of bacterial sensors in a very rational way, and it works quite well."
In its simulations and experiments, the lab mutated the histidine kinase protein, which detects an input and releases a phosphate group that binds to the response regulator. That, in turn, binds to DNA to activate gene expression.
"When you turn on gene expression, that's your signal," Tabor said. "What's cool about these pathways is that when the input is absent, the sensor histidine kinase acts as a phosphatase instead and removes the phosphate group. When that happens, the response regulator falls off the DNA and no longer activates the gene expression."
He said that mutating the histidine kinase to favor turning the signal on or off can be used to control the actions of response regulators, and thus the amount of desired proteins produced.
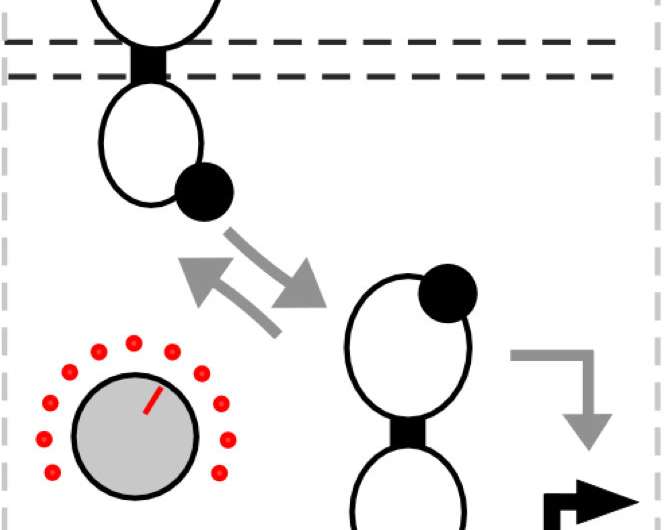
"We reasoned it's like a tug of war between the kinase activity and the phosphatase activity," Tabor said. "That balance determines how much input is needed to turn the sensor on. So this is like a knob where we can make the phosphatase activity stronger or weaker to tune the input."
He said the generality of the process should boost synthetic biology in general. "There are alternative methods to do what we did here, but they are far more labor intensive," Tabor said. "They're more likely to fail, and they'd take a whole Ph.D. to get them to work, whereas we can do this in a week and make it work."
More information: Brian P. Landry et al. Phosphatase activity tunes two-component system sensor detection threshold, Nature Communications (2018). DOI: 10.1038/s41467-018-03929-y
Journal information: Nature Communications
Provided by Rice University