Researchers establish crystal structure of gastric proton pump
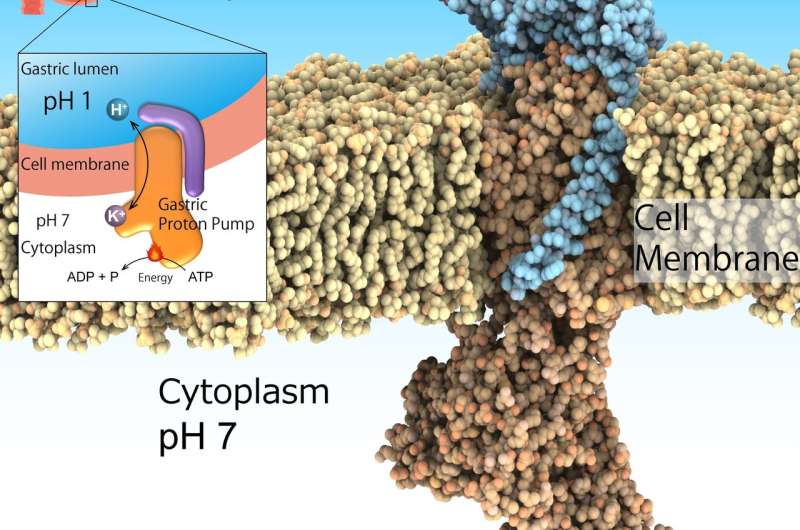
The highly acidic environment in the stomach is essential for digestion. Furthermore, it acts as an important barrier to invasive pathogens. However, excessive stomach acidification leads to ulcers. Although this is not a life-threatening condition, it can considerably impair the health of affected individuals. Acid suppression in combination with antibiotics is the recognized treatment to eradicate the bacteria Helicobacter pylori, a risk factor for gastric cancer. This environment of pH1 is regulated by the gastric H+, K+-ATPase, a class of enzymes that catalyze H+ transport from neutral cytosolic solution (pH 7) to the acidic gastric lumen (pH 1) fueled by cellular energy source ATP. Hence, gastric H+, K+-ATPase are prominent target for drugs that treat excess stomach acid secretion.
The main research question in this area of study is how such a highly acidic environment can be attained in the stomach. To address this question, the researchers sought the structure of H+, K+-ATPase. The team recently published their findings in Nature.
"In our study, we used X-ray crystallography to determine the structures of gastric H+, K+-ATPase bound to two proton pump inhibitors, vonoprazan and SCH28080," first author Kazuhiro Abe explains. "This information is important for both the refinement of existing drugs and the discovery of new drugs."
X-ray crystallography is a technique that uses X-ray diffraction patterns to determine high-resolution, three-dimensional structures of molecules such as proteins, small organic molecules, and materials.
The team successfully solved the crystal structure of H+, K+-ATPase in complex with either vonoprazan or SCH28080 to a resolution of 2.8 Å—high enough to reveal that the drugs partially overlapped but had clearly distinct binding modes in their binding sites found in the middle of a conduit running from the gastric lumen to the cation-binding site.
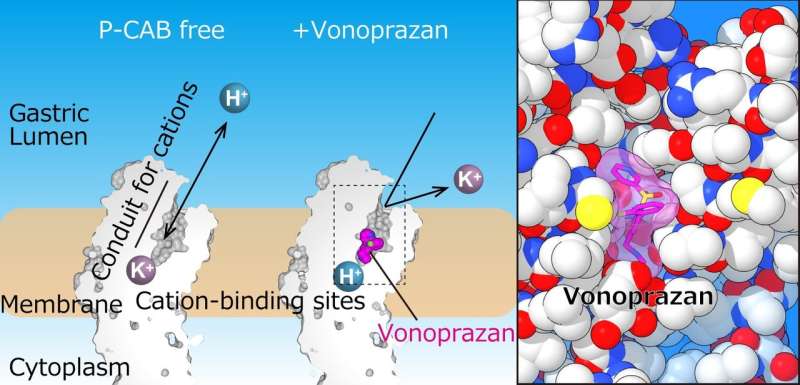
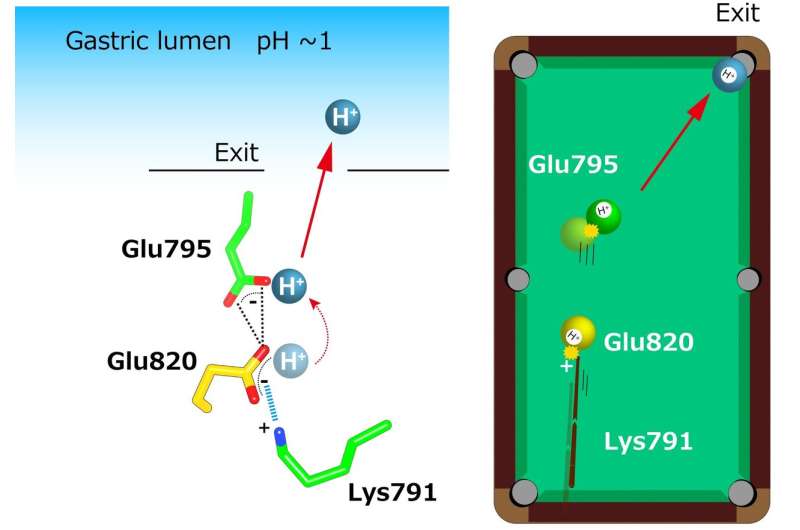
"The crystal structures suggest that the tight configuration at the cation-binding site lowers the pKa (a measure of acid strength) value of glutamic acid at residue 820 sufficiently to enable the release of a proton even into the pH 1 environment of the stomach," senior author Yoshinori Fujiyoshi says. "These structures define the molecular interaction between P-CABs (K+-competitive acid blockers) and H+, K+-ATPase, and reveal how H+, K+-ATPase expels H+ into the stomach even at pH1. Such information will contribute significantly to the knowledge base for drug discovery for conditions related to excessive stomach acidification."
More information: Kazuhiro Abe et al, Crystal structures of the gastric proton pump, Nature (2018). DOI: 10.1038/s41586-018-0003-8
Journal information: Nature
Provided by Nagoya University