Nanodiscs catch misfolding proteins red-handed
When proteins misfold, accumulate and clump around insulin-producing cells in the pancreas, they kill cells. Now, researchers, including University of Michigan biophysicists, have obtained a structural snapshot of these proteins when they are most toxic, detailing them down to the atomic level.
The researchers hope this kind of detail can help in the search for drugs to target the incorrectly folding proteins.
The clumps of misfolded proteins, called plaques or amyloid fibers, are implicated in many diseases. Amyloids interfere with neuron function in the brains of people with Alzheimer's and Parkinson's, and they also kill islet cells, which produce insulin to regulate blood sugar levels in people with type 2 diabetes.
"In general, toxicity to cells is extremely difficult to prove and characterize," said lead researcher Ayyalusamy Ramamoorthy, U-M professor of biophysics and chemistry. "On the other hand, we need to do this in order to help develop drugs for potential treatment."
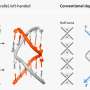
To understand the protein structure, the researchers use nanodiscs composed of layers of lipids surrounded by a belt—they look like a miniscule sushi roll. These lipids, bound by the belt, captures the proteins during their aggregation. The researchers then allow the protein to fold to a certain point within the nanodisc—when they think the folding proteins are most toxic to islet cells.
"The nanodiscs are like the difference between a swimming pool and the ocean. In the ocean, there are no boundaries; a swimming pool has boundaries," Ramamoorthy said.
"We're able to stop the aggregation of the protein in this restricted membrane environment so we can monitor what it looks like before it becomes a mass of fibers."
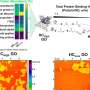
At this point, the researchers use a technique called nuclear magnetic resonance spectroscopy, or NMR, to make atomic-level images of the proteins. Just as an MRI scan takes images of the body so that physicians can diagnose an illness, NMR images proteins so researchers can study how they might be malfunctioning.
The researchers, who include U-M chemists and biophysicists Diana Rodriguez Carmago and Kyle Korshavn and others from the Technical University of Munich and the Helmholtz-Zentrum Muenchen, also hope to use the technique to both develop and screen for drug compounds that can target the misfolding proteins that are implicated in aging-related diseases including Alzheimer's disease and prion disease.
"We are screening small molecule compounds to see if we can inhibit this aggregation process that produces amyloids," Ramamoorthy said. "This has been much wanted—and much awaited information—for the scientific understanding of the pathology of amyloid diseases, and for the development of compounds to overcome the problems."
The researchers have been developing these "sushi-like" nanodiscs to get snapshots of these attacking proteins and characterize them for various biological and biomedical applications. These nanodiscs are also used to study other proteins in the cell membrane and how different proteins interact with each other in the cell membrane. The ability to pin down proteins while they're in the process of amyloid aggregation allows researchers to characterize the proteins using a variety of biophysical tools.
More information: Diana C Rodriguez Camargo et al. Stabilization and structural analysis of a membrane-associated hIAPP aggregation intermediate, eLife (2017). DOI: 10.7554/eLife.31226
Journal information: eLife
Provided by University of Michigan