Researcher discusses the biological crosstalk between microbes and hosts
Stavroula Hatzios listens in on dialogues between infectious bacteria and host cells. Trained in chemistry at MIT and Berkeley, she joined West Campus in January 2017 as a faculty member of the Microbial Sciences Institute and the Department of Molecular, Cellular, and Developmental Biology. Hatzios became interested in infectious disease as a graduate student and decided to focus on microbiology as a Harvard postdoc. Today her lab applies chemical tools to study how dangerous microbes, including the one that causes cholera, interact with the host. What she learns could point the way toward new strategies to fight infection.
What do you mean by host-microbe interactions?
There are many layers of communication that can exist between microbial and host cells in an infected animal. I refer to it as a dialogue. Microbes can use proteins and small molecules to communicate with one another as they form large communities, and similarly, host cells can secrete molecules that allow them to communicate with each other. And sometimes this crosstalk can also facilitate communication between host cells and microbial cells.
What kinds of dialogues are you hearing?
We study the cholera bacterium, Vibrio cholerae. Let's say you're in Haiti following the earthquake in 2010, when there was a breach of the water supply, and you ingest water that has cholera bacteria in it. The bacteria can end up colonizing your small intestine, and once they're there, they can produce a number of different proteins that can impact how the host responds to the infection and how the bacteria survive. One of those proteins is cholera toxin, one of the main mediators of the diarrhea that develops in infected hosts. That leads to massive dehydration, and the loss of fluid can kill quite rapidly. But the bacteria also produce other proteins that can be important for their interactions with the host, or their survival.
To query that dialogue between host and microbes, we applied a technique called activity-based protein profiling. This approach uses small-molecule probes that can bind to specific proteins based on their biochemical activity. We were able to identify a bunch of different bacterial- and host-secreted proteins with defined biochemical activities in animal models of cholera and in human choleric stool.
How many proteins are we talking about?
In this case, we identified over 200. For many of them we don't know what they do. Step one is to identify these proteins that are active in the infection. Step two is figuring out what they're doing. And step three is using them as tools–either to inhibit the development of disease by inhibiting their biochemical activity, or using them as diagnostic markers, or biomarkers, for disease.
During my postdoc, we identified these four bacterial proteases, which are enzymes that can chop up other proteins. These are produced by the cholera bacterium in the intestine–they were consistently identified in every single one of our analyses of infected rabbits. We were able to connect them to a host protein that's secreted by rabbit intestinal cells and binds to the cholera pathogen's cell surface. We found that these four pathogen-secreted enzymes inhibit the binding of this protein to the bacterial surface.
The finding that this interaction occurs in an infected animal–and that the cholera pathogen can produce proteases to inhibit this interaction–is really interesting. We believe this interaction may extend to other intestinal microbes, as well. Several other established groups are now investigating this possibility, and I'm excited to see what they discover. Is this interaction somehow regulating the composition of microbial communities in the intestine? Is it regulating how intestinal pathogens interact with the host? Is it helping to clear the infection, or are the bacteria using it to stick to host cells in some manner?
It's like analyzing warfare.
It is! One of the proteases that we identified in this work with cholera seems to be active only in an infected animal. Why that's interesting is that you can imagine ways to exploit the activity of that enzyme for your own benefit. One way we are hoping to do that is by generating Trojan horse-type probes—proteins engineered to mimic natural protease substrates, but which carry hidden cargo. The idea is that you could introduce these probes to the site of an infection, and they would only become activated when a specific pathogen-secreted protease is present and active. That would enable the spatially and temporally controlled release of, let's say, detection agents, or bactericidal agents.
Where is your research headed next?
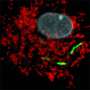
We think that reactive oxygen species–or small-molecule oxidants that are produced during infections–can help serve as a means of communication between host cells and neighboring microbes. Historically, these oxidants are considered cytotoxic (toxic to cells). But in the last two decades or so, there's been a lot of beautiful work showing that low levels of these oxidants are produced by host cells that mediate broader aspects of development, like differentiation, wound healing, motility. We're interested in how they influence signaling in microbial cells, or in host cells, once they are generated following microbial contact with the host.
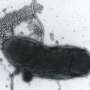
This oxidative crosstalk work that we do uses Helicobacter pylori, a gastric pathogen that causes stomach cancer. I am excited to hopefully collaborate with our new colleague here, Jun Liu. His lab is also interested in Helicobacter, and they've done some fascinating work looking at the flagella of that bacterium using a technique called cryo-electron tomography.
Provided by Yale University