Bioengineers discover mechanism that regulates cells' 'powerhouses'
UCLA bioengineers and their colleagues have discovered a new perspective on how cells regulate the sizes of mitochondria, the parts of cells that provide energy, by cutting them into smaller units.
The researchers wrote that this finding, demonstrated with yeast proteins, could eventually be used to help address human diseases associated with an imbalanced regulation of mitochondria size—for example, Alzheimer's or Parkinson's diseases. In addition, since having mitochondria that are too small or too large can potentially lead to incurable diseases, it is conceivable that the proteins responsible for this process could be potential targets for future therapies.
The study was published in ACS Central Science and was led by UCLA bioengineering professor Gerard Wong.
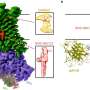
Inside the cell, mitochondria resemble the long balloons used to create balloon animals. If the mitochondria are too long, they can get tangled. Their sizes are known to be primarily regulated by two proteins, one of which breaks up longer mitochondria into smaller sizes. They are known as cells' "powerhouses" as they convert chemical energy from food into a form useful for cells to perform all their functions.
Keeping mitochondria at optimal sizes is important to cells' health. An insufficient amount of the regulating protein, known as Dnm1, results in the mitochondria getting too long and tangled. Too much Dnm1 results in too many short mitochondria. In both cases, the mitochondria are rendered essentially ineffective as power providers for the cell. This situation could lead to neurodevelopmental disorders or neurodegenerative diseases, such as Alzheimer's or Parkinson's.
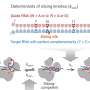
To better understand this mechanism, the researchers used a machine-learning approach they developed in 2016 to figure out exactly how the proteins break up one mitrochondrion into two smaller ones. They also used a powerful technique called "synchrotron small-angle X-ray scattering" at the Stanford Synchrotron Radiation Lightsource, a U.S. Department of Energy research facility, to see how these proteins deform mitochondrial membranes during this process.
Before this study, it was thought that these proteins encircled the mitochondria, then cut it in two by simply squeezing tightly. The process, the team discovered, is more subtle.
"When Dnm1 wraps around mitochondria, it has been previously shown that the protein physically tightens and pinches," said Michelle Lee, a recent UCLA bioengineering doctoral graduate who was advised by Wong and is one of two lead authors of the study. "What we found is that when Dnm1 contacts the mitochondrial surface, it also makes that area of the mitochondrion itself more moldable and easier to undergo cleavage. These two effects work hand in hand to make the process of mitochondrial division efficient."
The other lead author is Ernest Lee, a graduate student in the UCLA-Caltech Medical Scientist Training Program and a bioengineering graduate student also advised by Wong. He carried out the computational analyses for the experiment.
"Using our machine-learning tool, we were able to discover hidden membrane-remodeling activity in Dnm1, consistent with our X-ray studies," Lee said. "Interestingly, by analyzing distant relatives of Dnm1, we found that the protein gradually evolved this ability over time."
"This is a very unexpected result—no one thought these molecules would have a split personality, with both personalities necessary for the biological function," said Wong, who is also a UCLA professor of chemistry and biochemistry and is a member of the California NanoSystems Institute. "The multifunctional behavior we identified may be the rule rather than the exception for proteins."
More information: Michelle W. Lee et al. Molecular Motor Dnm1 Synergistically Induces Membrane Curvature To Facilitate Mitochondrial Fission, ACS Central Science (2017). DOI: 10.1021/acscentsci.7b00338
Journal information: ACS Central Science
Provided by University of California, Los Angeles