New algorithm expands use of advanced camera for biological microscopy

A new computer algorithm allows scientists to use a high-performance sensor technology, called scientific complementary metal-oxide semiconductor cameras, for a wide range of biological research.
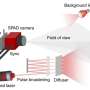
"Scientific sCMOS cameras are rapidly gaining popularity in biological sciences, material sciences and astronomy," said Fang Huang, an assistant professor in Purdue University's Weldon School of Biomedical Engineering. "The sensor provides significant advances in imaging speed, sensitivity and field of view compared with traditional detectors such as charge-coupled devices or electron multiplying CCD."
However, the use of sCMOS cameras for biological research has been limited because of fluctuations in pixel quality, generating more "noise," than the other cameras. In particular, every pixel fluctuates at its own rate.
"When you are trying to use this for biological studies, it's very difficult to determine whether this fluctuation comes from the sample (photons) or from the camera itself," said Sheng Liu, the lead author of the paper, a postdoctoral research associate at the Weldon School of Biomedical Engineering.
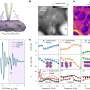
Now, working with researchers in Purdue's Department of Biological Sciences, Liu and Huang have developed a new algorithm that corrects the noise, making the sCMOS cameras available for a wide range of biological microscopy.
Findings were detailed in a research paper appearing earlier this year in the journal Nature Methods.
The authors include Liu; postdoctoral research associate Michael J. Mlodzianoski; graduate students Zhenhua Hu, Yuan Ren and Kristi McElmurry; Daniel M Suter, an associate professor in the Department of Biological Sciences; and Huang.
"We have been trying to use this camera for live-cell single-molecule super-resolution imaging and introduced an algorithm for that purpose in 2013," Huang said. "However, the previous algorithm works only for single-molecule studies, which means all your objects have to be so-called point emitters. So, basically, your images must look like stars in the sky."
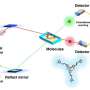
Biological research, however, often involves imaging complex structures such as cellular organelles. To solve the problem, researchers developed the new algorithm.
"The fundamental challenge is to estimate one of the variables when you know the sum of two variables. There's no unique answer to this question but we want to make the best estimate given our additional knowledge of the two variables." Huang said. "We exploited a general property of imaging systems, the optical transfer function. Based on our knowledge of how each of the 4 million pixels on our camera chip behave, we are able to estimate the actual photon level at each pixel location. This is very exciting for us because this allows CMOS sensors to be used in a broad spectrum of imaging methods for quantitative biomedical and biological studies, improving their sensitivity, field of view and imaging speed."
More information: Sheng Liu et al. sCMOS noise-correction algorithm for microscopy images, Nature Methods (2017). DOI: 10.1038/nmeth.4379
Journal information: Nature Methods
Provided by Purdue University