Gauging evolutionary adaptation- are our models right?
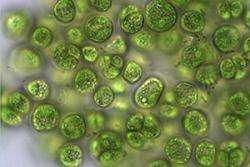
One challenge facing scientists is to estimate how our environment and the complex web of creatures within it, will respond to changes in their environment due to climate change or other human influences.
Traditionally, scientists have taken and tested single or pairs of ecological 'drivers' of change in the environment, elements such as increased temperature, increased CO2 or changes in herbicides or fertilizer, to assess how species will evolve over hundreds of generations.
This lab-based model of evolutionary change is simple compared to the complex environment in which species exist, so one major task for scientists is to understand how well simplified versions of environmental change teach us about more complex ones.
Writing in the latest online edition of PNAS evolutionary biologists at Edinburgh University and one now at Bangor University's School of Biological Sciences, tested how a single-celled alga evolved in environments with only a single driver versus in environments with many drivers, which are thought to be more representative of the multiple changes that organisms experience in the wider environment beyond the lab. The results suggest that the simplified environments with only a few drivers, provide a reasonably accurate idea of how much microbial populations evolve. This is because a few dominant drivers explain most of the evolutionary response of the population, even when other environmental drivers are present.
Using single-celled algae, which exists at the base of the food-chain, the study subjected the algae to 96 different environments with between 1 and 8 drivers, and allowed the algae to evolve in those environments for 450 generations, which corresponds to several seasons of growing in the environment.

What the researchers found was that, in most cases, a few dominant drivers (including temperature and CO2 levels) explained most of the evolutionary changes that took place.
They also found that despite the numerous variants tested, most populations converged on the same growth rates by the end of the 450 generations. However, environments with more drivers are initially more stressful and cause the populations to grow more slowly at the beginning of evolution, so the populations adapted more when the dominant drivers occur in the presence of other drivers – they simply had to evolve more to get to the same place.
Lead author Georgina Brennan, now at Bangor University's School of Biological Sciences explained:
"Aquatic primary producers will evolve under multiple changes, due to their rapid cell division rates and ability to generate genetic variation. Gauging the intensity of evolutionary selection within key primary producers at the base of the food chain will help understand how population sizes at the base of the food webs and nutrient cycles are impacted."
Sinéad Collins of Edinburgh University added:
"The power of simplified lab experiments lies in providing insights into how natural selection works, but applying the results of this experiment to natural phytoplankton population requires taking into account regional differences both in the phytoplankton populations and in the evolutionary drivers. I think that this is a hopeful study partly because it shows that evolution in simplified environments can give us insights into what happens in more complex environments – of course, it does mean that it is very important to correctly identify what the dominant drivers are in any given situation."
More information: Georgina L. Brennan et al. Evolutionary consequences of multidriver environmental change in an aquatic primary producer, Proceedings of the National Academy of Sciences (2017). DOI: 10.1073/pnas.1703375114
Journal information: Proceedings of the National Academy of Sciences
Provided by Bangor University