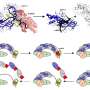
Solving a sweet problem for renewable biofuels and chemicals

Whether or not society shakes its addiction to oil and gasoline will depend on a number of profound environmental, geopolitical and societal factors.
But with current oil prices hovering around $50 dollars a barrel, it won't likely be anytime soon.
Despite several major national research initiatives, no one has been able to come up with the breakthrough renewable biofuel technology that would lead to a cheaper alternative to gasoline.
That research challenge led ASU scientists, Reed Cartwright and Xuan Wang, to enter the fray, teaming up to try to break through the innovation bottleneck for the renewable bioproduction of fuels and chemicals.
"My lab has been very interested in converting biomass such as agricultural wastes and even carbon dioxide into useful and renewable bio-based products," said Wang, an assistant professor in the School of Life Sciences. "As a microbiologist, I'm interested in manipulating microbes as biocatalysts to do a better job."
To do so, they've looked into a new approach—harnessing the trial-and-error power of evolution to coax nature into revealing the answer.
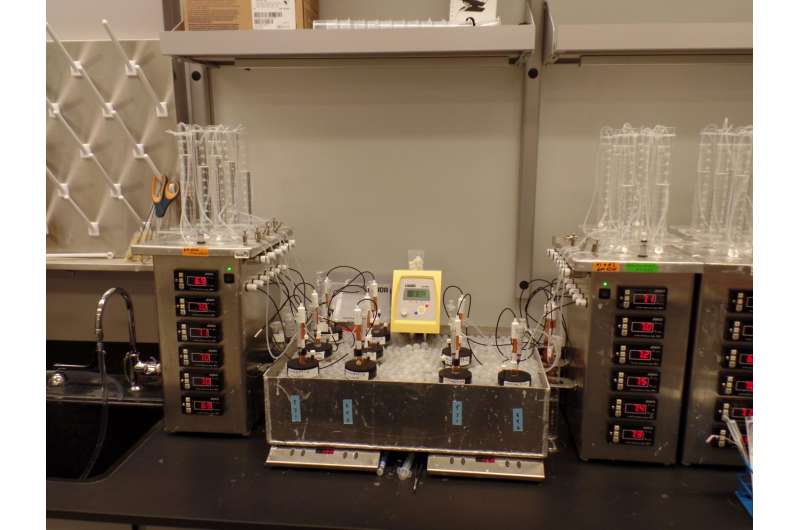
By growing bacteria over generations under specially controlled conditions in fermentation tanks, they have test-tube evolved bacteria to better ferment sugars derived from biomass—a rich, potential renewable energy source for the production of biofuels and chemicals.
Their results appeared recently in the online edition of PNAS.
The research team includes postdoctoral scholar Christian Sievert, Lizbeth Nieves, Larry Panyon, Taylor Loeffler and Chandler Morris, and was led by Reed Cartwright and Xuan Wang, in a collaboration between the ASU's School of Life Sciences and the Biodesign Institute.

A sweet problem
The appeal of plants is ideal. Just add a little carbon dioxide, water and plentiful sunshine, and presto! Society has a rich new source of renewable carbons to use.
Corn ethanol (using starch from corn for alcohol production primarily in the U.S.) has been one major biofuel avenue, and sugarcane another alternative (abundant in Brazil), but there is a big drawback. Turning the sugar-rich kernels of corn or sugarcane into ethanol competes with the food supply.
So scientists over the past few decades have migrated to research on conversion of non-food based plant materials into biofuels and chemicals. These so-called lignocellulosic biomasses, like tall switchgrasses and the inedible parts of corn and sugarcane (stovers, husks, bagasses, etc.) are rich in xylose, a five-carbon, energy-rich sugar relative of glucose.
Lignocellulosic biomass has an abundance of glucose and xylose, but industrial E coli strains can't use xylose because when glucose is available, it turns off the use of xylose. And so, to date, it's been an inefficient and costly to fully harvest and convert the xylose to biofuels.
Benchtop evolution
Wang and Cartwright wanted to squeeze out more energy from xylose sugars. To do so, they challenged E coli bacteria that could thrive comfortably on glucose—and switch out the growth medium broth to grow solely on xylose.
The bacteria would be forced to adapt to the new food supply or lose the growth competition.
They started with a single colony of bacteria that were genetically identical and ran three separate evolution experiments with xylose. At first, the bacteria grew very slowly. But remarkable, in no more than 150 generations, the bacteria adapted, and eventually, learned to thrive in the xylose broth.

Next, they isolated the DNA from the bacteria and used next-generation DNA sequencing technology to examine the changes within the bacteria genomes. When they read out the DNA data, they could identify the telltale signs of evolution in action, mutations.
Nature finds a way
The bacteria, when challenged, randomly mutated their DNA until it could adapt to the new conditions. They held on to the fittest mutations over generations until they became fixed beneficial mutations.
And in each case, when challenged with xylose, the bacteria could grow well. Their next task was to find out what these beneficial mutations were and how did they work. To grow better on xylose, the three bacterial E. coli lines had "discovered" a different set of mutations to the same genes. The single mutations the research team identified all could enhance xylose fermentation by changing bacterial sugar metabolism.
"This suggests that there are potentially multiple evolutionary solutions for the same problem, and a bacterium's genetic background may predetermine its evolutionary trajectories," said Cartwright, a researcher at ASU's Biodesign Institute and assistant professor in the School of Life Sciences.
The most interesting mutation happened in a regulatory protein called XylR whose normal function is to control xylose utilization. Just two amino acid switches in the XylR could enhance xylose utilization and release the glucose repression, even in the non-mutated original hosts.
Through some clever genetic tricks, when the XlyR mutant was placed back in a normal "wild-type" strain or an industrial E. coli biocatalyst, it could also now grow on xylose and glucose, vastly improving the yield. Wang's team saw up to a 50 percent increase in the product after 4 days of fermentation.
Together, Wang and Cartwright's invention has now significantly boosted the potential of industrial E. coli to be used for biofuel production from lignocellulosic materials. In addition, they could use this same genetic approach for other E. coli strains for different products.
Arizona Technology Enterprises (AzTE) is filing a non-provisional patent for their discovery. Wang hopes they can partner with industry to scale up their technology and see if this invention will increase economic viability for bioproduction.
"With these new results, I believe we've solved one big, persistent bottleneck in this field," concluded Wang.
More information: Christian Sievert et al, Experimental evolution reveals an effective avenue to release catabolite repression via mutations in XylR, Proceedings of the National Academy of Sciences (2017). DOI: 10.1073/pnas.1700345114
Journal information: Proceedings of the National Academy of Sciences
Provided by Arizona State University