May 2, 2017 report
Toward a better understanding of structure-metabolism relationships in human aldehyde oxidase (Update)
(Phys.org)—Drug design involves guided trial-and-error. How the body metabolizes a particular drug is important for determining drug efficacy. There have been many studies to understand how xenobiotics interact with cytochrome P450s, an important class of enzymes in drug metabolism, but little research has been done to understand aldehyde oxidase (AOX) metabolism. AOX, located in the liver, plays an important role in drug metabolism; however, many potential drugs end up failing during late-stage trials because of their interaction with AOX.
To better understand the structure-metabolism relationship between functional groups and AOX as well as to establish general guidelines for non-cytochrome P450 drug metabolism, researchers from the University of Perguia in Italy conducted tests on 198 compounds with aza-aromatic scaffolds to see which ones were oxidized by AOX. Furthermore, they tested 75 amide scaffolds to determine which ones were hydrolyzed. They found that electronic factors as well as steric hindrance affected the molecule's orientation in the MoCo active site, which determined whether the compound was metabolized by AOX. Their work appears in the Proceedings of the National Academy of Sciences.
AOX is an enzyme located in the liver that tends to oxidize aza-aromatic compounds as a phase I metabolite. Studies have shown that AOX activates the unsubstituted carbon ortho to the nitrogen on the aza heterocycle. Because this is the most electropositive carbon of the aromatic ring, electron density likely plays a role in AOX activation. But these studies, as well as others that look at AOX metabolism, were based on a small number of molecules that does not allow for deducing a general pattern for structure-metabolism relationship.
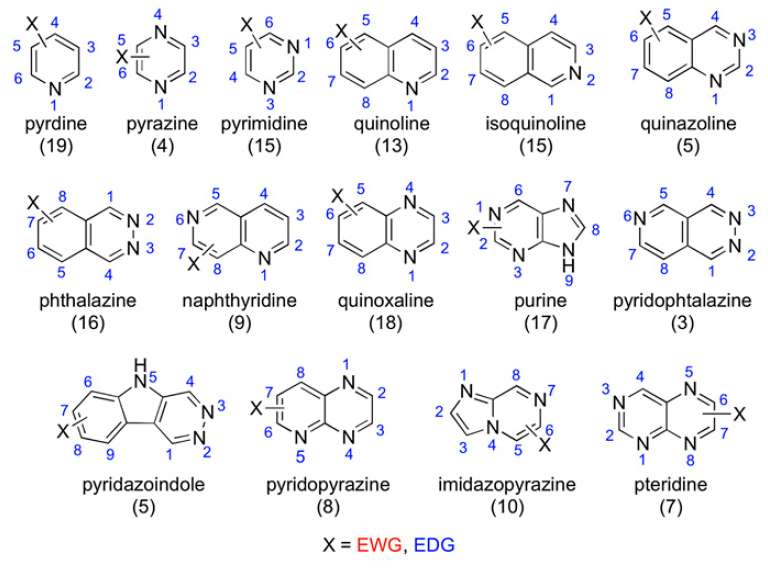
In the current study, Lepri et al. tested 198 aza-aromatic compounds to see if they were oxidized by AOX. Each of these had typical aza-aromatic scaffolds with variations on electron withdrawing and electron donating substituents. They either acquired or made this catalog of compounds and then conducted in vitro metabolism studies using human liver cytosol.
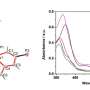
DFT calculations were used to elucidate the effects of electron density for AOX activity. The site of metabolism on the aromatic compound tended to correspond to the most electro-positive unsubstituted carbon, as expected. Additionally, computations studies were used to conduct docking analysis of the compounds in the active site of AOX.
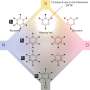
The authors found some trends for the site of metabolism; however, these trends are complicated by several factors. The pyridine scaffold was the only one that, as a scaffold class, was not susceptible to AOX metabolism. The other groups strongly depended on the electron density on particular carbons or, as the authors found with compounds such as quinoxalines and certain bicyclic compounds, steric hindrance in the active site prevented AOX metabolism.
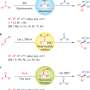
Lepri et al. also tested 73 amide compounds to see if AOX hydrolyzed the amide bond. In general, if there was an electron withdrawing group in the ortho position on the analine, then AOX did not oxidize it. If there was an electron donating group, then it did. The meta and para versions of electron donating and electron withdrawing groups were not susceptible to AOX metabolism.
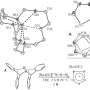
Exposure effects were an important component to whether a compound was metabolized by AOX. This has to do with the molecule's orientation toward the MoCo center in the AOX active site. Certain bulky groups resulted in no activity where one was expected based on computational studies. Additionally, hydrophobicity also affected how the compound interacted with the active site. When the reactive carbon is exposed to the MoCo center, then the compound is easily metabolized. But, when the unreactive portion of the compound is oriented toward the MoCo center, then the compound is less likely to be oxidized or hydrolyzed.
The authors point out that from these experiments "it emerges that it is not at all simple to predict whether a compound is a substrate of AOX or not." The difficulty lies in understanding the reactivity of the electrophilic carbon on the compound as well as how that particular compound will orient itself in the enzyme's active site. However, this study provides a starting point for additional studies and more sophisticated modeling techniques.
More information: Susan Lepri et al. Structure–metabolism relationships inAOX: Chemical insights from a large database of aza-aromatic and amide compounds, Proceedings of the National Academy of Sciences (2017). DOI: 10.1073/pnas.1618881114
Abstract
Aldehyde oxidase (AOX) is a metabolic enzyme catalyzing the oxidation of aldehyde and aza-aromatic compounds and the hydrolysis of amides, moieties frequently shared by the majority of drugs. Despite its key role in human metabolism, to date only fragmentary information about the chemical features responsible for AOX susceptibility are reported and only "very local" structure–metabolism relationships based on a small number of similar compounds have been developed. This study reports a more comprehensive coverage of the chemical space of structures with a high risk of AOX phase I metabolism in humans. More than 270 compounds were studied to identify the site of metabolism and the metabolite(s). Both electronic [supported by density functional theory (DFT) calculations] and exposure effects were considered when rationalizing the structure–metabolism relationship.
Journal information: Proceedings of the National Academy of Sciences
© 2017 Phys.org