What bone proteomics could reveal about the dead
Studying bones has helped scientists reconstruct what dinosaurs and other extinct creatures looked like. Taking this further, scientists recently started identifying proteins from bones to glean more information about remains. But one team has found that the reliability of this approach can depend on which bone is analyzed. Additionally, they report in ACS' Journal of Proteome Research a forensic use for bone proteomics: potentially determining from bone proteins how old someone was when they died.
Coaxing a picture of life from a person's or animal's bone proteins has been a tricky proposition. There have been few studies on how the profiles of proteins change as bones grow, whether the protein sets are the same across bones from the same individual, or even whether different parts of the same bone produce similar groups of proteins. At the same time, interest among archeologists, paleontologists and forensic scientists in using bone proteomics for determining species relationships has grown, in addition to improving our understanding of the fossilization process. So Noemi Procopio, Michael Buckley and colleagues set out to explore the types of information that proteins from bony remains might reveal.
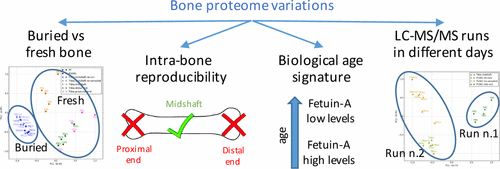
To find out, the researchers extracted proteins from either fresh or buried pig bones. Testing revealed that different parts of an individual animal's skeleton yielded different sets of proteins. As expected, the most reproducible results came from proteins extracted from the midshaft of long bones, such as the tibia, or shin bone, at positions farthest from the growth plates. The study also identified some proteins that either decreased or increased in samples with increasing biological age. The researchers say such information could potentially act as biomarkers for age in human remains, if further studies bear out this finding.
More information: Noemi Procopio et al. Intra- and Interskeletal Proteome Variations in Fresh and Buried Bones, Journal of Proteome Research (2017). DOI: 10.1021/acs.jproteome.6b01070
Abstract
Proteomic methods are acquiring greater importance in archaeology and palaeontology due to the longevity of proteins in skeletal remains. There are also developing interests in forensic applications, offering the potential to shed light on post-mortem intervals and age at death estimation. However, our understanding of intra- and interskeletal proteome variations is currently severely limited. Here, we evaluated the proteomes obtained from five distinct subsamples of different skeletal elements from buried pig carcasses to ascertain the extent of variation within and between individuals. We found that reproducibility of data depends on the skeletal element used for sampling and that intrabone differences exceed those observed between the same skeletal element sampled from different individuals. Interestingly, the abundance of several serum proteins appeared to correlate with biological age with relative concentrations of alpha-1 antitrypsin and chromogranin-A increasing and those of fetuin-A decreasing. We also observed a surprising level of divergence in data from different LC–MS/MS runs on aliquots of similar samples analyzed months apart, adding constraints to the comparison of results of such methods across different studies.
Journal information: Journal of Proteome Research
Provided by American Chemical Society

















