Automated evaluation speeds up the search for new active substance
The macromolecular crystallography beamlines at the BESSY II X-ray source are specially designed to highly automated structural analyses of protein crystals. With up to now more than 2,000 solved structures of protein molecules, these beamlines are by far the most productive ones in Germany and are in vigorous demand by groups from either an academic and industrial research area.
Now teams from HZB and Philipps-Universität Marburg in Germany have automated the evaluation of data records as well. The newly developed expert system identifies small molecule fragments bound to proteins in the raw X-ray diffraction data. These fragments represent suitable starting points for the development of an active substance. Using a series of 364 samples, the collaborating partners demonstrated that the expert system works reliably and can speed up the search for a suitable active agents.
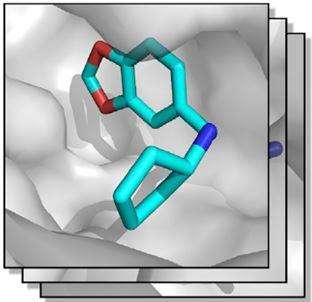
The search for new substances that are effective against a disease is like the first step in cracking a high-security lock: the active substance needs to fit into the target molecule like a key in a lock. The target molecule is in this case a protein, responsible for a specific function in an organism. The degree to which this function is accomplished can be influenced by active agents. Thus, an appropriate active substance can correct a function that has gone out of control and causes a disease. However, you first need to identify such an appropriate substance. And this search can be very tedious. Figuratively speaking, an infinite number of keys can be imagined, and it would take a huge amount of time to test them all. An efficient safecracker will not spend a lot of time checking such a large bunch of keys, instead he will study the pins of the lock, pin after pin, until he can set each pin correctly. A similar procedure, called fragment screening, became the conventional approach in structural biology. To discover a prototypal "blueprint", you assemble several individual fragments to a larger entity. To do so, you test hundreds of very small molecules (fragments) as to whether they bind to the target protein. A first lead structure for an active substance is then assembled from those fragments that bind.
High troughput and strong demand: the MX-Beamlines
A highly automated version of this procedure is already running at the BESSY II MX beamlines for three years and facilitates high throughput of samples. With over 2000 solved protein structures so far, the macromolecular crystallography beamlines at BESSY are by far the most productive ones in Germany. They are heavily overbooked with demands originating from academic as well as industrial research user groups.
New expert system evaluates raw data
Now a collaborative research project between the Philipps-Universität Marburg and HZB has succeeded in bringing this automation a step further. "We are able to work through hundreds of samples using fragment screening where manual evaluation is hardly feasible any longer", says Dr. Manfred Weiss, who heads the HZB MX team. In collaboration with Prof. Gerhard Klebe and his group at Marburg, the teams jointly developed an expert system as a computer program that incorporates the expertise of scientists. This expert system evaluates the raw X-ray crystallography data using an iterative process pipeline of evaluation routines and identifies those fragments that bind to the protein. The group headed by Klebe made the protein crystals available together with a library of different fragment molecules suitable for protein binding. In collaboration, the partners tested the newly developed expert system using crystals of a specific protein soaked in total with 364 fragments in aqueous solutions.
Automatic refinement identifies additional candidates
The expert system is also capable to automatically refine the X-ray structures. "Without this automated refinement procedure of the iterative expert system, a fair portion of the fragments would have remained undetected", explains Weiss. "For example, we had only identified about 50 candidates during the first evaluation, while by using the iterative adaptive procedure an additional 32 candidates could be discovered." The work shows that the new expert system allows a more reliable detection of fragments that can be considered as suitable starting points or promising molecular portions for the development of potent active agents. This can speed up the search for new drug molecules.
More information: Johannes Schiebel et al, High-Throughput Crystallography: Reliable and Efficient Identification of Fragment Hits, Structure (2016). DOI: 10.1016/j.str.2016.06.010
Journal information: Structure
Provided by Helmholtz Association of German Research Centres