Scientists find new system in tomato's defense against bacterial speck disease
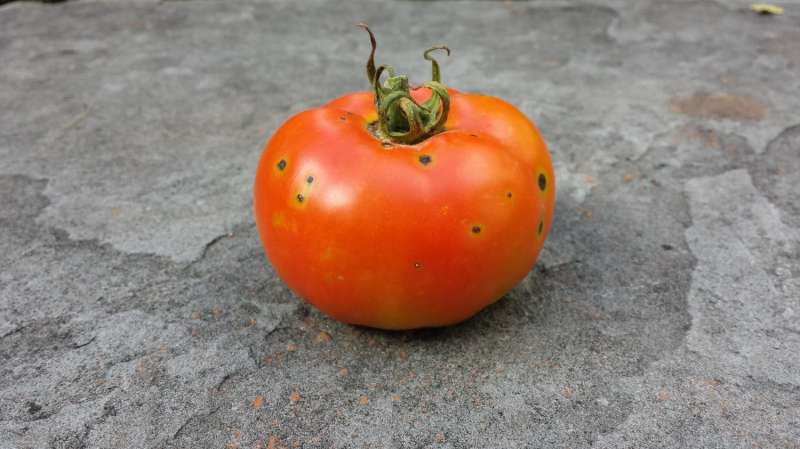

Researchers at the Boyce Thompson Institute (BTI) and Virginia Tech have discovered a new mechanism in the continual arms race between plants and pathogenic bacteria, which tomatoes use to detect the causal agent of bacterial speck disease.
The team identified a new receptor in tomato plants, called FLAGELLIN-SENSING 3 (FLS3), that triggers defenses against a bacterial attack. FLS3 is present in a small number of plant species, including tomato, potato and pepper. The study appears in Nature Plants.
"This is an interesting example of a receptor that appears to have evolved fairly recently because it is only found in a small group of plants," said first author Sarah Hind, a research associate in the lab of Professor Gregory Martin at BTI. "This discovery sets up the possibility of introducing FLS3 into other economically important crop plants, which might provide resistance to bacterial pathogens that is not naturally present in other plants."
FLS3 detects a part of the flagellum, a tail-like appendage that helps bacteria swim through their environment and consists mostly of flagellin proteins. When bacteria invade the plant, the FLS3 receptor binds to a region of the flagellin protein called flgII-28 and triggers an immune response.
FLS3 is the second flagellin sensor discovered in tomatoes. The first, called FLS2, is found in most land plants and detects invading bacteria by recognizing a separate region of flagellin called flg22. Since many plants have an FLS2 receptor, several bacterial species have acquired mutations in the flg22 portion of their flagellin, and these mutations alter the flagellin shape so that FLS2 can no longer recognize it. The acquisition of the FLS3 receptor may therefore serve as a countermeasure on behalf of the tomato to detect bacteria that have altered flg22.
Martin and his colleagues worked with postdoctoral researcher Christopher Clarke and Professor Boris Vinatzer at Virginia Tech, who previously identified flgII-28 as a conserved segment of the flagellin protein that tomatoes can detect. Together, the researchers demonstrated that flgII-28 could not be recognized by FLS2, suggesting the existence of the yet unknown FLS3. Additionally, in collaboration with Associate Professor Dilip Panthee of North Carolina State University, the researchers screened heirloom tomato varieties and found that some, including Yellow Pear, could not respond to flgII-28, suggesting that the tomato must be missing FLS3.
"Discovering that some heirloom tomatoes, such as Yellow Pear, did not respond to flgII-28 was key to using a genetics approach to identify FLS3," said senior author Martin, the Boyce Schulze Downey Professor at BTI and a Cornell University professor in the School of Integrative Plant Science.
In the current paper, Hind used Yellow Pear tomatoes together with a wild relative of tomato called Solanum pimpinellifolium to identify the FLS3 gene and show how it functions to reduce bacterial growth. But to confirm that FLS3 is the receptor for flgII-28, she needed to demonstrate that the two molecules can physically interact. Researchers in the laboratories of Martin and BTI Associate Professor Frank Schroeder developed cutting-edge biochemistry techniques to identify a stable complex between FLS3 and flgII-28, thus validating FLS3 as the flgII-28 receptor.
"Proving direct interactions of biomolecules has remained a huge challenge, and our work will help in developing better approaches for exploring receptor-ligand interactions," said co-author Schroeder,a Cornell University professor in Chemistry and Chemical Biology.
The study demonstrates how versatile the plant immune system can be while fighting a constant arms race against infectious bacteria. "Plants are always coming up with new ways to defeat pathogens," said Hind. "We're trying to understand how they do it and then use this knowledge to develop more disease-resistant plants."
More information: Sarah R. Hind et al, Tomato receptor FLAGELLIN-SENSING 3 binds flgII-28 and activates the plant immune system, Nature Plants (2016). DOI: 10.1038/nplants.2016.128
Journal information: Nature Plants
Provided by Boyce Thompson Institute