Powerful new metric quickly reveals network structure at multiple scales
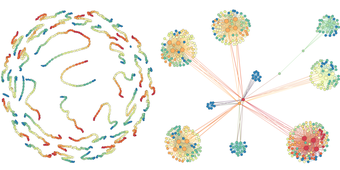
What does a network look like? It typically depends on what scale you're analyzing.
Researchers often want to know what hidden structures lie within data representing real-world networks, from power grids to the internet. To do this, they employ a variety of methods and metrics, but these methods are limited. Approaches that identify microscopic features miss the big structural picture. Methods that reveal macroscopic organization don't reliably show how the network is constructed – and also tend to be computationally intensive.
"We don't have a good toolbox to get a quick understanding of the network structure," says Laurent Hébert-Dufresne, a James S. McDonnell Fellow at the Santa Fe Institute. The recent explosion in data available to researchers, both in the variety of sources and in the sheer size of the datasets, emphasizes the need for improved methods to analyze and synthesize structural information about complex networks.
Hébert-Dufresne's frustration with the current state of network analysis inspired him – together with Antoine Allard of the University of Barcelona and Joshua Grochow, an SFI Omidyar Fellow – to devise a new tool. The trio recently developed a new metric that hopscotches over the limitations of other methods, revealing network structure at the microscopic, mesoscopic, and macroscopic levels in one clean blow. They introduced their approach, which they dubbed the "Onion Decomposition" as a metaphor for peeling back the layers of an onion, this week in Scientific Reports.
Complex networks are usually depicted as nodes, or dots, connected by edges, or lines. The researchers' new tool analyzes a network by taking it apart – peeling away "layers" of nodes that have the same number of connections, usually starting with the layers having the fewest numbers of connections. That's not new, strictly speaking; the same approach is used by another powerful algorithm called "k-core decomposition."
However, as it peels away the layers, k-core decomposition loses valuable information about those layers, says Hébert-Dufresne. His group's goal was to build an algorithm that capitalizes on the benefits of k-core decomposition but also uses layer-level information to provide insights about the network at multiple scales.
In their paper, the researchers report successful test drives of their method on a handful of real-world datasets, including the Northwestern U.S. power grid and the road system of Pennsylvania. In both cases, the onion delivered accurate snapshots of the network structures at different scales, and the authors were able to draw some interesting conclusions. By removing a few nodes in the power grid network, for example, the network's overall connectivity would quickly collapse; the road network is more robust, however, and the network would remain more or less well connected despite the removal of a few nodes.
Hébert-Dufresne sees the new metric as a valuable first step for network analysis that "allows us to understand, at a glance, whether a network is tree-like or grid-like, how heterogeneous it is, and even identify surprising subgraphs."
For instance, the paper demonstrates the easy detection of long chains of websites that do nothing but hold hands in subsets of the world wide web. The method could improve models of disease spread by not only considering how connected individuals are, but also how central they are (that is, how deep they are in the layers of a network).
Since creating the tool, he and his collaborators have been using it in their own research in a variety of settings, from social networks to food webs.
"The best testament for why we think this will be useful, even now that the project is done, is that this is a tool that we use," says Hébert-Dufresne.
More information: Laurent Hébert-Dufresne et al. Multi-scale structure and topological anomaly detection via a new network statistic: The onion decomposition, Scientific Reports (2016). DOI: 10.1038/srep31708
Journal information: Scientific Reports
Provided by Santa Fe Institute