Amino acid sequences are key to the properties of silks
Although mankind has been using silk for millennia, many aspects of its properties and molecular structure remain unknown. Now, a new study from the RIKEN Center for Sustainable Resource Science (CSRS) in Japan reveals that amino acid sequences are key determinants of the material properties of silk fibers. The work was performed by CSRS scientists in an international collaboration with Beihang University, Indian Institute of Technology Guwahati and Chulalongkorn University.
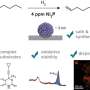
The study, published in Scientific Reports, aimed to shed light on the determinants of the unique properties of silkworm silks. Silks fibers are made up of large structural proteins, called fibroins, featuring highly repetitive amino acid sequence modules. However, there has been considerable debate as to whether the amino acid sequence or the fiber spinning process itself plays the critical role in determining material properties.
To probe these questions, the authors compared silks obtained from a wide variety of silkmoths; silk is produced not just by the larvae of the domesticated silkmoth, Bombyx mori, but also by a diverse array of "wild" silkmoths, including species that have so far received little attention. The genes encoding silk fibroins have evolved over hundreds of millions of years, and different groups of silkmoths have adopted different strategies for producing silk fibers that optimize their survival.
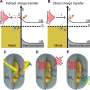
One unique aspect of the study was that the different silk samples were subjected to a battery of different materials tests, including tensile deformation, thermal analysis, and characterization of the crystalline component of the silk fibers, the latter being carried out at the SPring-8 synchrotron facility in Harima.
The results provided several new insights into the properties of silk. For instance, the differences in the repetitive sequence arrangement among the different types of silk helps explain why, in domesticated silk, stretching of the fibers produces relatively soft or gradual structural transitions, whereas in the wild silks these are observed as sharp or concerted transitions. Furthermore, among the wild silks, a clear correlation was shown between the length of the crystalline-forming regions of the fibroin sequence and thermal stability. Overall, the authors conclude that repetitive amino acid sequences of different silk types play a decisive role in determining their unique physical properties.
According to Keiji Numata, leader of the Enzyme Research Team at CSRS, "The results of this study have major implications for future efforts to design and produce artificial silk proteins. Although silk is best known for its use in textiles, recently there has been an explosion of interest from fields such as biomedical research and materials science, due to the rare combination of strength, extensibility, biocompatibility a biodegradability exhibited by silk." Ali Malay, lead author of the study, further commented: "Various attempts have been made to replicate the material properties of native silks using recombinant bacterial expression or via chemical synthesis routes, but these efforts have mostly yielded disappointing results. Understanding the intimate connections between amino acid sequence, molecular structure, and material properties would enable the fine-tuning of silk-like sequences to achieve specific desired material properties."
More information: Scientific Reports, DOI: 10.1038/srep27573
Journal information: Scientific Reports
Provided by RIKEN