RNA simulations boost understanding of retroviral diseases
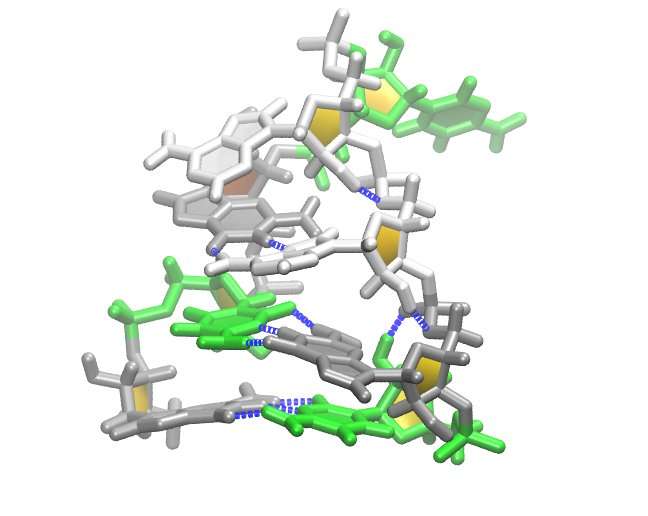
New molecular dynamics research into how RNA folds into hairpin-shaped structures called tetraloops could provide important insights into new treatments for retroviral diseases.
"Ribonucleic acid, known as RNA, forms the genome of multiple viruses that afflict humans, including Ebola, HIV and Zika, which are active areas of research at Los Alamos," said Jacob Miner, a Los Alamos National Laboratory graduate student in the Center for Nonlinear Studies and a member of the research team. "Being able to describe the structures and thermodynamic behaviors of these and other RNA molecules is critical for us as scientists to understand life at microscopic levels and to effect suitable measures to treat diseases."
"One of the most exciting discoveries in our computer models is the emergence of a particular type of tetraloop that we thought only formed in highly saline environments," said Miner. "Our results show that similar sequences may be spontaneously forming these tetraloops, called Z-form duplexes, in nature, providing a potential target for treating retroviral diseases," he noted.
Research at Los Alamos has advanced the science of molecular biology over the years through such efforts as using the Laboratory's supercomputers to create the world's largest computational biology simulation of ribosomes in action. Another project, detecting pathogens through RNA screening, has recently been opened up for corporate partnering. In addition, Los Alamos scientists are teaming with other institutions in human trials of a complex mosaic approach to a globally applicable HIV vaccine. Biological research, especially in disease forecasting and treatment, aids in national security in preventing or minimizing the effect of epidemics and stabilizing the health of the nation.
For the new RNA research, Miner and a team from Rensselaer Polytechnic Institute and the State University of New York, Albany pursued a goal of the nucleic-acid modeling community, that of understanding the various structures that linear chains of RNA molecules can form. While DNA has the well-known "double helix" structure, RNA's single chain permits it to writhe and contort into completely different shapes, each giving the RNA a different function, such as coding and decoding genes or building proteins.
The prevalence of one RNA configuration over another is driven by its thermodynamic stability, and the relative populations of these configurations can be modulated by changes in the solution environment such as temperature, pressure, ions and the presence of other molecules.
Molecular dynamics (MD), which allows scientists to model detailed molecular interactions of RNA with itself and solution, has proven to be one of the most useful tools in understanding the modulation of RNA configurations, Miner said. "The behaviors exhibited in MD simulations can be extended to describe macroscale thermodynamic properties in all manner of RNA structures. In the case of our research, it represents a significant advancement in the utility of RNA molecular dynamics for describing RNA thermodynamics."
More information: Jacob C. Miner et al, Free-energy landscape of a hyperstable RNA tetraloop, Proceedings of the National Academy of Sciences (2016). DOI: 10.1073/pnas.1603154113
Journal information: Proceedings of the National Academy of Sciences
Provided by Los Alamos National Laboratory