April 26, 2016 report
Prion-like proteins found in mustard plant
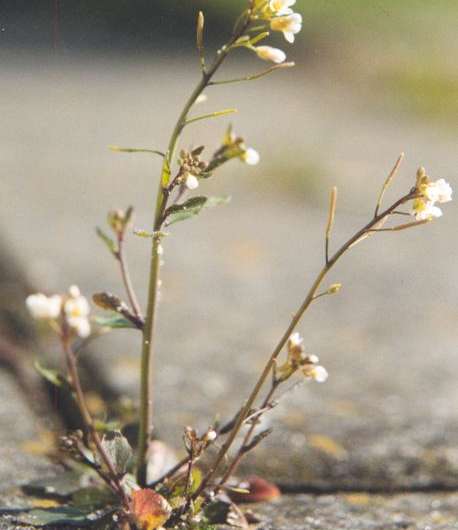

(Phys.org)—A team of researchers affiliated with the Whitehead Institute for Biomedical Research, MIT and the Howard Hughes Medical Institute, all in Massachusetts has found a prion-like protein in Arabidopsis thaliana, a flowering mustard plant. The group has published a paper describing their experiments and results in Proceedings of National Academy of Sciences.
Prions are most well known as being the cause of mad-cow disease, but they are also found in other species of animals as well, and now, new evidence suggests they may also appear in some plants.
Prions are a type of protein that fold under certain conditions and when they do, they cause other proteins around them to fold as well, which is how they lead to neurological damage in cows. In this new effort, the research team, which is more often concerned with studying properties of yeast, began to wonder if prions might be involved in plant memory—prior research had shown that prions are able to serve a memory function in yeast.
To find out, they ran an algorithm that sifted through all of the proteins that have been identified in A. thaliana, which uncovered approximately 500 possible candidates—they narrowed that list down to just four through closer analysis, each of which are involved in flowering. This was important, the team notes, because mustard plants have demonstrated a type of memory—holding off flowering, for example, during cold snaps, which are not true indicators of the onset of winter.
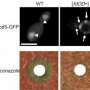
The researchers then cut the Sup35 protein from a yeast sample (it has been identified as the part that is involved in prion folding), and replaced it with the protein from the mustard plant and then watched how the proteins behaved. The team reports that one of the modified proteins did indeed change shape in a prion-like way, suggesting the possibility that it could actually be a prion. If so, it would be the first ever found in a plant.
In order to prove that the protein is in fact a prion, the plant would have to be taken apart and examined to see if the protein could be found to fold in its native environment, a challenge the team suggests that would best be tackled by plant specialists.
More information: Sohini Chakrabortee et al. Luminidependens (LD) is an Arabidopsis protein with prion behavior, Proceedings of the National Academy of Sciences (2016). DOI: 10.1073/pnas.1604478113
Abstract
Prion proteins provide a unique mode of biochemical memory through self-perpetuating changes in protein conformation and function. They have been studied in fungi and mammals, but not yet identified in plants. Using a computational model, we identified candidate prion domains (PrDs) in nearly 500 plant proteins. Plant flowering is of particular interest with respect to biological memory, because its regulation involves remembering and integrating previously experienced environmental conditions. We investigated the prion-forming capacity of three prion candidates involved in flowering using a yeast model, where prion attributes are well defined and readily tested. In yeast, prions heritably change protein functions by templating monomers into higher-order assemblies. For most yeast prions, the capacity to convert into a prion resides in a distinct prion domain. Thus, new prion-forming domains can be identified by functional complementation of a known prion domain. The prion-like domains (PrDs) of all three of the tested proteins formed higher-order oligomers. Uniquely, the Luminidependens PrD (LDPrD) fully replaced the prion-domain functions of a well-characterized yeast prion, Sup35. Our results suggest that prion-like conformational switches are evolutionarily conserved and might function in a wide variety of normal biological processes.
Journal information: Proceedings of the National Academy of Sciences
© 2016 Phys.org