Physicists develop new software for the life sciences

Scientists from Heidelberg University have developed a novel mathematical model to explore cellular processes: with the corresponding software, they now are able to simulate how large collections of cells behave on given geometrical structures. The software supports the evaluation of microscope-based observations of cell behaviour on micropatterned substrates. One example is a model for wound healing in which skin cells are required to fill a gap. Other areas of application lie in high throughput screening for medicine when a decision needs to be taken automatically on whether a certain active substance changes cell behaviour. Prof. Dr. Ulrich Schwarz and Dr. Philipp Albert work both at the Institute for Theoretical Physics and at the Bioquant Centre of Heidelberg University. Their findings were recently published in PLOS Computational Biology.
One of the most important foundations of the modern Life Sciences is being able to cultivate cells outside the body and to observe them with optical microscopes. In this way, cellular processes can be analysed in much more quantitative detail than in the body. However, at the same time a problem arises. "Anyone who has ever observed biological cells under a microscope knows how unpredictable their behaviour can be. When they are on a traditional culture dish they lack 'orientation', unlike in their natural environment in the body. That is why, regarding certain research issues, it is difficult to derive any regularities from their shape and movement," explains Prof. Schwarz. In order to learn more about the natural behaviour of cells, the researchers therefore resort to methods from materials science. The substrate for microscopic study is structured in such a way that it normalises cell behaviour. The Heidelberg physicists explain that with certain printing techniques, proteins are deposited on the substrate in geometrically well-defined areas. The cell behaviour can then be observed and evaluated with the usual microscopy techniques.
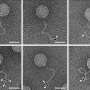
The group of Ulrich Schwarz aims at describing in mathematical terms the behaviour of biological cells on micropatterned substrates. Such models should make it possible to quantitatively predict cell behaviour for a wide range of experimental setups. For that purpose, Philipp Albert has developed a complicated computer programme which considers the essential properties of individual cells and their interaction. It can also predict how large collections of cells behave on the given geometric structures. He explains: "Surprising new patterns often emerge from the interplay of several cells, such as streams, swirls and bridges. As in physical systems, e.g. fluids, the whole is here more than the sum of its parts. Our software package can calculate such behaviour very rapidly." Dr Albert's computer simulations show, for example, how skin cell ensembles can overcome gaps in a wound model up to about 200 micrometres.
Another promising application of these advances is investigated by Dr. Holger Erfle and his research group at the BioQuant Centre, namely high throughput screening of cells. Robot-controlled equipment is used to carry out automatic pharmacological or genetic tests with many different active substances. They are, for example, designed to identify new medications against viruses or for cancer treatment. The new software now enables the scientists to predict what geometries are best suited for a certain cell type. The software can also show the significance of changes in cell behaviour observed under the microscope.
More information: Philipp J. Albert et al. Dynamics of Cell Ensembles on Adhesive Micropatterns: Bridging the Gap between Single Cell Spreading and Collective Cell Migration, PLOS Computational Biology (2016). DOI: 10.1371/journal.pcbi.1004863
Journal information: PLoS Computational Biology
Provided by Heidelberg University