Seeing cancer cells in 3-D (w/ Video)
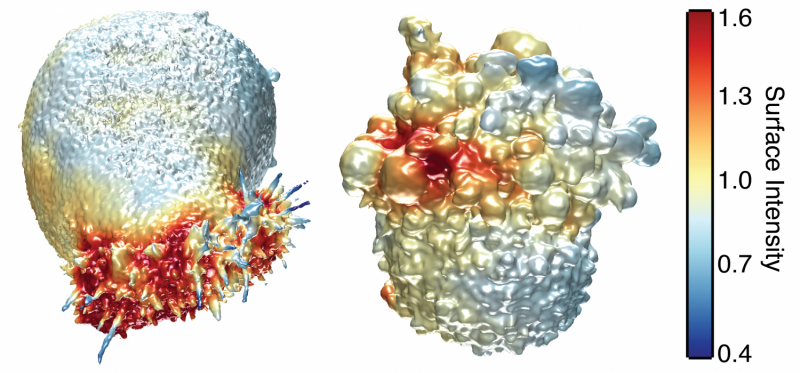
Cancer cells don't live on glass slides, yet the vast majority of images related to cancer biology come from the cells being photographed on flat, two-dimensional surfaces—images that are sometimes used to make conclusions about the behaviour of cells that normally reside in a more complex environment. But a new high-resolution microscope, presented February 22 in Developmental Cell, now makes it possible to visualize cancer cells in 3D and record how they are signaling to other parts of their environment, revealing previously unappreciated biology of how cancer cells survive and disperse within living things.
"There is clear evidence that the environment strongly affects cellular behavior—thus, the value of cell culture experiments on glass must at least be questioned," says senior author Reto Fiolka, an optical scientist at the University of Texas Southwestern Medical Center. "Our microscope is one tool that may bring us a deeper understanding of the molecular mechanisms that drive cancer cell behavior, since it enables high-resolution imaging in more realistic tumor environments."
In their study, Fiolka and colleagues, including co-senior author Gaudenz Danuser, and co-first authors Meghan Driscoll and Erik Welf, also of UT Southwestern, used their microscope to image different kinds of skin cancer cells from patients. They found that in a 3D environment (where cells normally reside), unlike a glass slide, multiple melanoma cell lines and primary melanoma cells (from patients with varied genetic mutations) form many small protrusions called blebs. One hypothesis is that this blebbing may help the cancer cells survive or move around and could thus play a role in skin cancer cell invasiveness or drug resistance in patients.
The researchers say that this is a first step toward understanding 3D biology in tumor microenvironments. And since these kinds of images may be too complicated to interpret by the naked eye alone, the next step will be to develop powerful computer platforms to extract and process the information.
"When we conceived of this project, we first asked what we wanted to measure and then designed a microscope and analytical platform to achieve this goal," says co-first author Erik Welf, a cell biologist. "We hope that now instead of asking what we can measure, scientists will ask what we must measure in order to make meaningful contributions to cancer cell biology."
The microscope control software and image analytical code are freely available to the scientific community.
More information: Developmental Cell, Welf and Driscoll et al.: "Quantitative Multiscale Cell Imaging in Controlled 3D Microenvironments" dx.doi.org/10.1016/j.devcel.2016.01.022
Journal information: Developmental Cell
Provided by Cell Press