Fighting antibiotic resistance—how bacteria knit their 'sugar armour' at the single-molecule level
In a new paper published in Nature Chemistry, Dr. Lingbing Kong in Oxford University's Department of Chemistry takes an in-depth look at capsular polysaccharides, or 'sugar armour' – the outermost layer of bacteria that provides a key defensive shield for pathogens (including against antibiotics).
Here, Dr Kong describes the latest research:
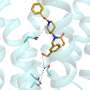
'I started the work contained in the paper in 2007 in the labs of Professor Ben Davis and Professor Hagan Bayley in Oxford's Department of Chemistry. The overall aim of my project was to elucidate the interaction of the capsular polysaccharide (CPS) K30 and its exporter outer membrane protein Wza, which would facilitate the development of novel antibacterial strategies. The biochemical, biophysical, and biological part of the project was going smoothly and led to the discovery of the first inhibitor of the Wza pore. It is, however, fundamentally important to analyse the interaction of the K30 CPS and the Wza pore at the single-molecule level, which would be crucial for later generations of the Wza inhibitor when resistant mutants appear.
'This part of the project had several barriers that had to be overcome – it has been a great challenge in academia to successfully analyse this kind of bimolecular interaction. The carbohydrate-protein interactions are generally weak, and the access to defined oligosaccharides and polysaccharides is always the main limiting factor. Therefore, we dedicated ourselves to achieving this goal.
'This new Nature Chemistry paper reports how we achieved this over the last nine years and reveals new insights into the important biological process.
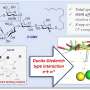
'We have developed a generalised synthetic approach to obtain polysaccharides with defined sizes, which has rarely been attempted in literature. In polysaccharide-related syntheses, it has been a frequent bias that only one of the possible repeating ("monomer") units, typically the most synthetically accessible, is selected for synthesis. Our approach includes a non-biased analysis of the targeted polysaccharide for all possible minimum oligosaccharide repeating units. Next, multi-step organic synthesis gave the desired repeating units, which were then activated to form the according oligomers and polymers.
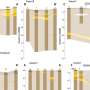
'Further separation by high-performance liquid chromatography afforded the defined large oligosaccharides and polysaccharides in pure forms. The analysis of the interaction between K30 carbohydrate fragments and the exporter Wza pore was carried out with an advanced droplet system that requires a volume as small as 200 nanolitres. This setup was developed to enable analysis and detection of the behaviours of the same protein pore before and after the addition of the sugar substrates, which was not possible previously. As a result, the weak interaction between K30 CPS and the Wza pores was detected at the single-molecule level.
'Analysis of the complex data revealed that only small (not large) fragments of the K30 fragments were translocated through the pore. We also observed capture events that occur only on the intracellular side of Wza, which would complement coordinated feeding by adjunct biosynthetic machinery.
'The techniques that we developed to recapitulate sugar export at the single-molecule level would potentially allow studies of all bacterial sugar export. This will facilitate not only the detection of the sugar-exporter interactions but also screening of blockers of the sugar exporters. In addition, the generalised approach that we developed for polysaccharide synthesis, ie polyglycosylation, could potentially be applied to all polysaccharide synthesis. Such approaches are rarely reported and usually not generalisable.
'Our approaches exemplified: 1) non-biased analysis of the chemical structures of the target polysaccharide; 2) rational design and solid synthesis of all polymerisable minimum oligosaccharide repeating units; 3) screening and optimization of conditions for polyglycosylation; and eventually 4) separation of individual oligomeric and polymeric products, which is followed by deprotection to afford defined and pure large oligosaccharides and polysaccharides.
'On the other hand, the advanced droplet system that we developed enables the detection and analysis of weak interactions between a membrane protein pore and the substrates in a very small quantity at the single-molecule level. For one single experiment, only one protein pore and 1 microgram (or even 1 nanogram) of substrate (given molecular weight 500 and a concentration of 1 minimole/litre or 1 micromole/litre, respectively) are required.
'Nowadays, mass spectrometry is one of the most sensitive techniques. The technology we developed would enable the detection and analysis of bimolecular interactions in the same resolution. We therefore expect our technologies to find far-reaching applications in numerous vital binding processes in glycobiology.'
More information: Lingbing Kong et al. Chemical polyglycosylation and nanolitre detection enables single-molecule recapitulation of bacterial sugar export, Nature Chemistry (2016). DOI: 10.1038/nchem.2487
Journal information: Nature Chemistry
Provided by University of Oxford