Scientists blueprint antimicrobial candidate that may stem post-antibiotic tide
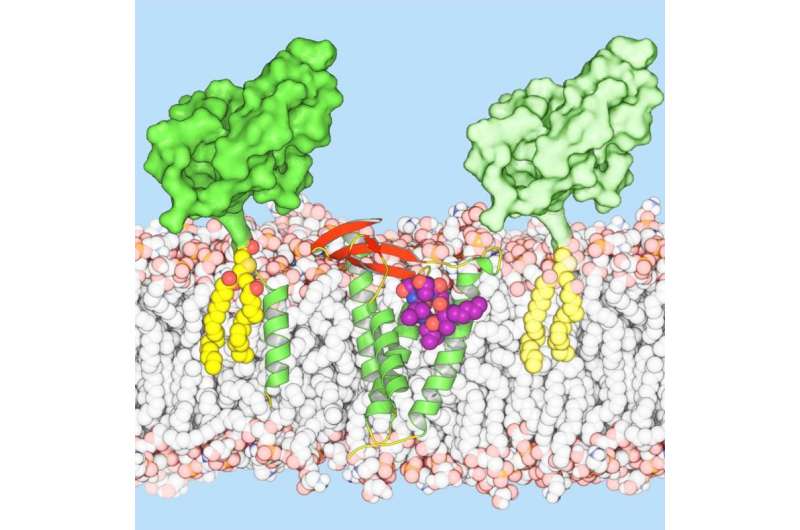
Scientists at Trinity College Dublin have provided the first crystal clear molecular blueprint of Globomycin - an antibacterial candidate with promise in stemming the onrushing post-antibiotic tide.
Crucially, their blueprint may aid the design of better globomycin analogues and the exploration of thousands of other potential new antibiotic solutions to common but devastating infections.
With antimicrobial resistance on the rise, the World Health Organization has advised that a post-antibiotic era, in which minor injuries and common infections could prove fatal, is looming. New drugs are badly needed.
Globomycin attracted attention as an antibacterial candidate of significant promise more than a decade ago, because it inhibits a key enzyme LspA and thus the production of many cell wall proteins with critical roles in bacterial physiology, pathogenicity and antibiotic resistance. If delivered effectively, Globomycin and Globomycin-like drugs could knock out a variety of bacterial infections.
However, the progression of earlier studies stagnated, partly because researchers did not possess the all-important high-res blueprints with which to guide further drug development. That hurdle has now been removed as that very blueprint has just been published in a landmark paper in leading international journal Science.
Professor of Membrane Structural and Functional Biology at Trinity College Dublin, Martin Caffrey, said: "Our LspA-globomycin blueprint can be used to inform the design of globomycin derivatives or analogues with improved binding, efficacy, selectivity and pharmacokinetic properties."
"The molecular form of the complex makes it a suitable target with which to explore the thousands of inhibitors developed for other medically relevant aspartyl protease enzymes. These include drugs that treat high blood pressure and HIV AIDS."
The blueprint might also be used to design entirely new drugs targeted specifically at the chemical constellation of the enzyme's active site in the bacteria Pseudomonas aeruginosa. However, it may be that blueprints of LspA from other pathogenic bacterial species such as Enterococcus, Staphylococcus (MRSA), Klebsiella, Acinetobacter and Enterobacter should be the immediate focus of efforts at the design of species-specific antibiotic drugs.
Such so-called narrow spectrum antimicrobials are more attractive because they are less likely to promote a generalised antibiotic resistance response.
Professor Caffrey added: "What is especially exciting is that the blueprint makes it hard to conceive bacteria developing resistance to Globomycin or newly developed analogues."
"Globomycin and its enzyme target are networked to such a degree that any mutation perturbing the drug's interaction would also likely impact on how the bacteria functions. Antibiotics that do not elicit resistance are invaluable to medicine, and this work might inspire the design of such drugs."
In a twist of evolutionary fate, Globomycin is a natural antimicrobial produced by bacteria of the Actinomycete type, which means we are effectively drawing up battle plans to wage war against bacterial infections by borrowing from the arsenal of other bacteria. Lessons in developing effective drugs should be learned from this reality.
More information: Structural basis of lipoprotein signal peptidase II action and inhibition by the antibiotic globomycin," Science, DOI: 10.1126/science.aad3747
Journal information: Science
Provided by Trinity College Dublin