Innovative machine could accelerate discoveries in structural biology, drug design

Alexander Barnes and his colleagues are in the midst of a protracted and laborious experiment whose goal to discover how one biological molecule interacts with another. They recently measured a distance of about 0.00000002 inch between two atoms in their target. Admittedly, that's a tiny distance but it took 918,000 scans and 42 days of instrument time to gauge it. What could possibly justify this kind of time?
Designing a drug that will flush the HIV virus out of its hiding places within cells for one thing. When HIV enters cells, it makes DNA copies of its RNA core that it slips into the cell's chromosomes. Once that happens, antiretroviral drugs can prevent virus in the bloodstream from copying itself and infecting other cells but not clear the virus. The viral DNA persists in the cells, waiting to act as a template for new virus if the drugs are ever stopped.
This is why doctors are reluctant to say that AIDS is ever cured, even when there has been no detectable virus in the blood for some time.
Barnes, PhD, assistant professor in the Department of Chemistry in Arts & Sciences, and colleagues at Washington University in St. Louis and at Stanford University, hope to make a drug that will dock on cells and trick the hidden DNA into turning on and making and releasing RNA, which can then be destroyed by antiretroviral drugs. With its cellular machinery tied up working for the virus, the cell would not be able to maintain itself and would die.
"But to do this we need to know exactly how this drug is structured and how it moves at room temperature, and we don't know either of those right now," Barnes said. "We don't know what the structure is when its bound to the protein at room temperature. We just don't know.
"And we want to know so badly that we're taking eight months on an NMR spectrometer to figure it out."
Determining the three-dimensional structure of biological molecules such as proteins has always been a challenge. The first protein to be studied at atomic resolution was hemoglobin. Its structure was solved by X-ray crystallography in 1959. NMR, or nuclear magnetic resonance, began to contribute to structural biology in about 1985 but so far most structures have been determined by crystallography.
Both techniques allow scientists to learn the structures of molecules too small to be seen with the unaided eye. But both have major drawbacks: X-ray crystallography because proteins don't want to crystallize and it can take years to coax one to do so; and NMR because the signal intensities are so small that extensive signal averaging is required to get a signal to pop out of the noise.
Barnes' new machine uses a technique called dynamic nuclear polarization that theoretically could boost the signal by a factor of 600, allowing measurements to be completed 400,000 times faster. "Even if we can only manage to boost the signal by a factor of 100, we could do the experiment that took 42 days in 42 minutes," Barnes said.
"If we can determine the structures and motion of biomolecules 100 times faster than we do now, there's going to be huge application of this technology," he said. "It can be applied to study virtually every drug and every biomolecule of interest to science. Everyone will want to use it."
Why the signal is so weak
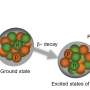
Subatomic particles such as electrons, protons and neutrons can have a quantum mechanical property called spin, and it is this property NMR spectroscopy detects. In many NMR samples, the spins are paired against one another and the net spin is close to zero. The tiny spins behave like tiny compass needles when they are exposed to an external magnetic field.
As physicists put it, they have a magnetic moment, which means they can be pushed by a magnetic field. If there is an external field they will try to align with it, or directly against it. Their spin axes trace out little circles centered on the field lines, rather like spinning tops that are about to topple over. This motion, called precession, generates an electromagnetic field with a characteristic frequency.
If the sample is irradiated with radio waves at the same "resonant" frequency, the nuclei will absorb the energy. When the radio frequency pulse ends, the nuclei will precess and generate an electrical signal at their characteristic frequency; it is this signal that is detected and analyzed to identify target nuclei.
So far so good, but here is the rub: In NMR, the energy separation between nuclear spin states is very small, small enough that the electrical signal is barely detectable.
Goosing the signal
"We want to do these experiments faster, much faster," Barnes said. To goose the nuclear spin polarization, his machine steals polarization from electrons.
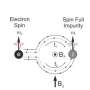
If a soap bubble filled with oxygen is put in a magnetic field, it will be drawn to the magnet. The reason is oxygen, as a consequence of the way it is put together, has two unpaired electrons, that is electrons existing as unpaired spins.
Crucially, because electrons have a tiny mass compared to protons they have much bigger magnetic moments. Six hundred times bigger, said Barnes, which also means that electrons can have a much bigger population difference between states.
By adding a "polarizing agent," a chemical that has many unpaired electrons, to the sample, the scientists can create a much bigger "spin reservoir."
The trick is to transfer the highly populated spin polarization of the electron to the less populated nuclear one. This is done by illuminating the sample with electromagnetic radiation at the electron's spin resonance, the frequency that will cause electrons to flip spin states. When this transition is saturated, polarization is transferred to nuclei in the sample.

The idea that the highly populated spin polarization of the electron could be transferred to the less populated nuclear one is not obvious and took some time to be accepted. In addition, there was a practical problem.
To get good resolution, the scientists needed to work with strong magnetic fields but this moves the spin resonance frequency of the electrons to high frequencies that are difficult to produce. One of the recent breakthroughs in the field is the development of high-power gigahertz (one billion cycles per second) microwave sources.
Barnes built one of these devices, called a gyrotron, as part of his doctoral work at MIT.
Coming in from the cold
But as things currently stand, dynamic nuclear polarization only works at cryogenic temperatures. It fails at room temperature, Barnes said, because it takes longer for the polarization to transfer from the electrons to the nuclei than for the electrons to relax back to their equilibrium states.
This is a problem, Barnes said, because biomolecules may be trapped in a configuration as they freeze that they would not assume at room temperature. And in any case, scientists are as interested in dynamics—how the molecules move over time—as in structure at any given moment.
"What we want to do is to sweep the microwave source through the electron resonance frequency," he said. "If we can sweep the frequency, the electrons feel it differently. One way to think of it is you are kind of pushing them over, pushing them from spin up to spin down. Whereas if you don't have a tunable source, if you can't sweep the frequency, you're just hitting them with electromagnetic radiation and it doesn't work that well."
Barnes just received a prestigious award from the National Institutes of Health that will help him to develop this new technology, called frequency swept dynamic nuclear polarization.
Should everything turn out as he hopes and fast NMR spectroscopy become a reality at room temperature, it might change structural biology as dramatically as the famous PCR reaction changed genetics.
Significantly in the acknowledgments for his doctoral dissertation, he thanks his father for inspiring him to become a scientist by "writing out polymerase chain reactions on napkins over dinner when I was in grade school."
Provided by Washington University in St. Louis