NMR 'fingerprinting' for monoclonal antibodies
National Institute of Standards and Technology researchers at the Institute for Bioscience and Biotechnology Research have demonstrated the most precise method yet to measure the structural configuration of monoclonal antibodies (mAbs), an important factor in determining the safety and efficacy of these biomolecules as medicines. Monoclonal antibodies are proteins manufactured in the laboratory that can target specific disease cells or antigens (proteins that trigger an immune reaction) for removal from the body. The method described in a recent paper may soon help manufacturers and regulators better assess and compare the performance and quality of mAbs.
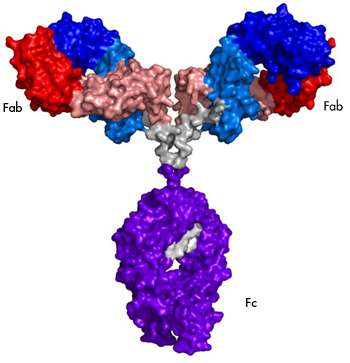
Monoclonal antibodies can be used as extremely specific therapeutic agents, including ones designed to target cancer cells unique to an individual. However, in order to properly function as a biotherapeutic agent, the molecule's structural units—amino acids—must fold into a three-dimensional structure that aligns its active regions with corresponding receptor sites on a target cell or antigen. If misfolding occurs, a potent and safe treatment may become ineffective, or worse, provoke a dangerous or fatal immune reaction. High-resolution spectral analysis—imaging at the atomic level where even the bonds between hydrogen and carbon atoms are distinguishable—is required to precisely define the mAb's structure and determine if the protein is folding properly.
"We refer to this as 'measuring fingerprints,' because just as a person has a unique set of fingerprint patterns, each mAb has a one-of-a-kind spectral makeup," says NIST research chemist Robert Brinson. "If we can map that spectral fingerprint, we can determine whether or not folding is occurring as desired."
To do this, the IBBR team turned to a solution that would surprise most biopharmaceutical experts: two-dimensional nuclear magnetic resonance (2D NMR) spectroscopy. NMR is a technique that measures the atomic signature of a molecule similar to how doctors use magnetic resonance imaging (MRI) to noninvasively view organs. "To date, it's been assumed that 2D NMR could not be practically applied to monoclonal antibodies because it's too insensitive, too time intensive and too expensive for analyzing anything other than much smaller drug molecules," Brinson explains.
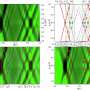
In pushing the boundaries of the technique, the IBBR team used an NMR system with a high magnetic field strength to produce the first 2D NMR map of a complete, drug-like mAb. The map was generated using signals from methyl groups.
"Methyl groups are dispersed throughout the mAb structure and, in particular, in the folded cores of the molecule that we want to evaluate," Brinson says. "We can use their signals to yield a specific spectral fingerprint that reflects the unique structure of the mAb."
To make the 2D NMR method more accessible to the lower-strength magnetic field instruments found in most analytical research labs, the IBBR team narrowed the analysis by dividing its sample antibody into two structural fragments."We mapped the 2D NMR signals generated by the subset of methyl groups found in these fragments, both about a third of the size of the entire protein," Brinson says. "The sum of the data gained from this analysis was found to be a good proxy for the spectral fingerprint of the full mAb."
The new 2D NMR fingerprinting method also overcomes the problems of cost and time. "We reduced the time needed for our measurements from many hours to about 30 minutes," Brinson says.
Brinson says that he and his colleagues are now working on a statistical method that will allow users of their 2D NMR methodology to compare fingerprints from multiple protein samples. "With that ability, manufacturers will be able to quantitatively show that spectra obtained from different lots of the same drug product are identical, enabling them to better meet regulatory requirements for quality and performance," he says.
More information: "Mapping monoclonal antibody structure by 2D 13C NMR at natural abundance." Analytical Chemistry, 87: 3556-3561 (2015). DOI: 10.1021/ac504804m
Journal information: Analytical Chemistry
Provided by National Institute of Standards and Technology