Researchers use neutron scattering and supercomputing to study shape of a protein involved in cancer
For living organisms proteins are an essential part of their body system and are needed to thrive. In recent years, a certain class of proteins has challenged researchers' conventional notion that proteins have a static and well-defined structure.
It's thought that mutations in these proteins, known as intrinsically disordered proteins, are associated with neurodegenerative changes, cardiovascular disorders and diseases such as diabetes and cancer. A better understanding of these proteins could have major implications in the energy, environmental and health biosciences.
One intrinsically disordered protein, beta-catenin, is of particular interest to researchers at the Department of Energy's Oak Ridge National Laboratory, where they will be conducting neutron scattering and supercomputing studies to further uncover its role in cancer.
"This protein violates everything we know about proteins," says Arvind Ramanathan of ORNL's Computational Data Analytics Group. "It doesn't have a very stable secondary structure except for a small region which forms a kidney bean shape."
"What we proposed to do is to take a look at the protein using neutron scattering and combine the data with large scale simulation of this molecule utilizing supercomputing capabilities at ORNL," he says.
The joint research project started after ORNL, Georgetown and Howard Universities, Washington Veteran's Affairs Medical Center and MedStar Research Institute won a Clinical Translation Science Award. The award is funded by the National Center for Advancing Translation Science, one of the institutes at the National Institutes of Health.
The beta-catenin protein occurs naturally in the body and is active in a pathway that sends signals from outside a cell to the inside of a cell. One of its terminal ends responds to cancer, and researchers have observed negative effects during cancer if changes occur to this side.
"Mutations in this protein usually impact drug resistance, relapse of cancers and many other important functional cellular roles during cancer," Ramanathan says.
To gain insight and a better idea of what beta-catenin's shape is and how the shape changes under various circumstances, the researchers plan to use neutron scattering techniques on ORNL's Spallation Neutron Source's EQ-SANS instrument.
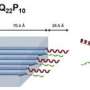
"On the neutron side, we are doing experiments that will allow us to characterize in very great detail how the protein changes its shape," Ramanathan says.
The data from the neutron experiments will then be analyzed using the supercomputing capabilities at DOE's Oak Ridge Leadership Computing Facility at ORNL. Additionally, the researchers will run simulations and model the protein while looking at other proteins to detect and characterize the protein's cancer responding terminal.
What they learn from the study will help them understand how the B-catenin protein binds to its partner cells and how the process changes after a mutation has occurred, which is often associated with cancer.
"If we don't understand how the protein binds specifically to certain signaling partners, we wouldn't know how to distinguish what its function is in a cancer related pathway versus a functional pathway," Ramanathan says.
Provided by Oak Ridge National Laboratory