March 26, 2015 report
New mitochondrially-derived peptides show what they can do
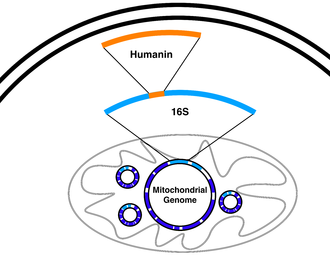
(Phys.org)—There is a whole lot more to the textbook mitochondrial genome then once was thought. A case in point is a multifunctional peptide named humanin that is dual-encoded deep within 16S ribosomal RNA gene in the mtDNA. Pinchas Cohen's lab was one of three labs that simultaneously co-discovered humanin when screening for proteins that may be involved in Alzheimer's, IGF-1 signaling, and apoptosis. Cohen's group just published a report in Cell Metabolism where they described another mitochondrially derived peptide, this time encoded within the 12S RNA-c gene, which has also has some useful properties. A mere 16 amino acids in length, they have demonstrated that this MOTS-c peptide as they call it (mitochondrial open reading frame of RNA-c) has dramatic effects on obesity and insulin resistance.
Before barreling ahead about the wonders of MOTS-c, we should probably say a bit more about humanin and mitochondrial translation in general. The final working length of humanin depends on where it was produced. If it was made inside the matrix using mitochondrial translation hardware its ends up being 21 amino acids long. If made outside mitochondria in the cytosol it will be 24 amino acids long. Both the tRNAs and rRNAs from mitochondria can end up getting transported into the cytosol, however, the mechanism and logic behind these processes is not yet understood.
In a previous post we noted that there are similarities between some nuclear encoded tRNAs (like met-tRNA) and the analogous met-mt-tRNAs, and therefore asked what it would take for mitochondrial to utilize nuclear tRNAs, and similarly vice-versa. The authors recognized the possibility that short peptides, like MOTS-c in particular, might be of nuclear origin due to the well-known phenomenon of nuclear mtDNA transfer (NUMT). After doing so-called BLAST searches they were not able to find any putative NUMT-derived peptides with complete homology to MOTS-c, but a search using human expression sequence tags (EST) found several hits for homologous mRNA sequences to the mitochondrial 12S rRNA locus.
This is a somewhat confusing state of affairs to say the least. Rats, for example, don't have any NUMT sequences for MOTS-c, making mtDNA its exclusive source. For the case of humanin (which in rats the homologous peptide is called, as you may have anticipated, 'rattin' ), other research has turned up at least 10 nuclear-encoded 'humanin-like' peptides expressed in human tissues. For short length peptides it may be difficult to conclusively say that these are cases of mitochondrial transfer, ie NUMTs, or just sequence convergence effects.
As for the question of where MOTS-c is made, its translation obligatorily occurs in the cytoplasm via the standard genetic code. This is because translation using the mitochondria-specific genetic code would yield tandem start and stop codons—hardly a useful message. The authors therefore reasonably suggest that the MOTS-c polyadenylated transcript is exported from the mitochondria. As for how MOTS-c is involved in obesity and diabetes, they point to altered gene expression of enzymes involved in the folate-methionine cycle and de novo purine synthesis. These results came from their microarray analysis and metabolomics profiling studies.
The full complexity of the mtDNA code, now unravelling before us, reflects the unique genetic systems of the mitrochondrion's bacterial ancestors. As we begin to comprehend the entire mitochondrial transcriptome, including its many small RNAs, we will be better equipped to evaluate potential fertilization outcome for issues like mitochondrial transfer in the creation of three parent embryos. Inclusive in any such discussion of effectively editing an entire mitochondrial genome in this blunt way would be consideration of the effects of new and contentious genome-editing techniques as they would be applied to mitohcondria themselves.
When we raised the question above about the potential for internexed mito- and nuclear- translation, we also noted recent research on the historically deep multi-read sequences of ribosomal subunits themselves which keynotes aspects of overlapping RNA codes. Although not fully vetted, nor rejected, it may be worthwhile to take a deeper look into that particular ribosome model, where the ribosome itself is a microcosm encoding a complete set of tRNS and accessory proteins, to better understand the origins of the larger genetic system we possess.
More information: The Mitochondrial-Derived Peptide MOTS-c Promotes Metabolic Homeostasis and Reduces Obesity and Insulin Resistance, Cell Metabolism, www.cell.com/article/S1550-413 … (15)00061-3/abstract
Abstract
Mitochondria are known to be functional organelles, but their role as a signaling unit is increasingly being appreciated. The identification of a short open reading frame (sORF) in the mitochondrial DNA (mtDNA) that encodes a signaling peptide, humanin, suggests the possible existence of additional sORFs in the mtDNA. Here we report a sORF within the mitochondrial 12S rRNA encoding a 16-amino-acid peptide named MOTS-c (mitochondrial open reading frame of the 12S rRNA-c) that regulates insulin sensitivity and metabolic homeostasis. Its primary target organ appears to be the skeletal muscle, and its cellular actions inhibit the folate cycle and its tethered de novo purine biosynthesis, leading to AMPK activation. MOTS-c treatment in mice prevented age-dependent and high-fat-diet-induced insulin resistance, as well as diet-induced obesity. These results suggest that mitochondria may actively regulate metabolic homeostasis at the cellular and organismal level via peptides encoded within their genome.
Journal information: Cell Metabolism
© 2015 Phys.org