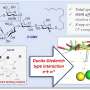
The promiscuity of chemical probes discovered

Researchers at IMIM (Hospital del Mar Medical Research Institute) have applied a new computational methodology to anticipate the degree of selectivity of the molecules that are used to study protein functions and reduce the risk of establishing erroneous relations between proteins and diseases. The proteins under study could be future candidates for new therapeutic targets. The study is published in the prestigious journal ACS Chemical Biology and was selected for the cover.
Molecules are essential tools for exploring protein functions, as they have the capacity to activate, inhibit and modulate their function. For many years, in order to explore protein functions, namely to know their biological role, small molecules known as 'chemical probes' have been used, which interact with the protein under study, to become a possible candidate as a new therapeutic target. However, in order for them to be truly useful, these molecules must selectively interact with the protein under study. 'Until now, it was assumed that these chemical probes only and exclusively interacted with the protein that was being studied, so that any variations in the results of experiments were interpreted as the consequence of the selective interaction of the chemical probe with the protein under study' comments Jordi Mestres, coordinator of the Research Group in Systems Pharmacology at the Research Programme on Biomedical Informatics (GRIB per its Spanish acronym) at IMIM and the UPF.
Through this research project, researchers have proven that many chemical probes are not selective. The truth is totally the opposite, as they interact with multiple proteins, often involved in the same biological pathways, which means that experimental results can be confused and lead researchers to draw mistaken conclusions with regard to the therapeutic relevance of many proteins. The consequences are truly important since, based on these erroneous conclusions, years and money can be invested in developing medicines that are ineffective and, above all, not safe.
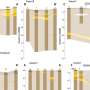
The project has consisted of computationally, and then experimentally, predicting that some 200 molecules from the National Institute of Health Program to Identify Chemical Probes in the United States have biologically relevant interactions with other proteins. To carry out this task, researchers have used a new computational method developed by the company Chemotargets SL, an IMIM spin-off created in 2006, specialising in developing software to design more efficient and safer drugs. These results have alerted researchers who use these molecules that they must bear the new interactions identified in mind when they interpret and draw conclusions on their experiments.
'Lack of knowledge about the interactions with other proteins can cause laboratories throughout the world to continue employing these "dirty" molecules to study a specific protein for many years. This entails an enormous loss of research time and resources', states Albert Antolín, a researcher in the same research group. 'This is why, before using molecules to study the biological function of a protein, its profile of interactions with proteins must be known as broadly and completely as possible, thus preventing researchers from drawing erroneous conclusions about the therapeutic function and relevance of the protein under study' adds the researcher.
As incredible as it may seem, even today we still know very little about the functions that many of the proteins in the human body have and the impact that the loss of these functions have on human health. 'Correctly characterising protein functions is therefore key to discovering new more efficient and safer medicines' conclude the researchers.
More information: "Distant Polypharmacology among MLP Chemical Probes", Albert A. Antolín and Jordi Mestres. ACS Chem Biol DOI: 10.1021/cb500393m
Journal information: ACS Chemical Biology
Provided by Hospital del Mar Medical Research Institute