Solving the Hox Specificity Paradox
The remarkable diversity of anatomical features along the body axis of animals—the differences between the head, the thorax and the abdomen, for example—is determined by proteins in the Hox family. But almost as soon as the Hox genes were discovered, scientists began puzzling over how different Hox proteins could activate specific genes, because all of these proteins can bind to the same DNA sequences. Now Howard Hughes Medical Institute scientist David Stern says he and his colleagues have solved the paradox.
Stern, a group leader at HHMI's Janelia Research Campus, and Janelia postdoctoral fellow Justin Crocker investigated the "Hox Specificity Paradox" in collaboration with Richard Mann at Columbia University. Their research shows that Hox proteins trigger gene activity through weak interactions at previously unrecognized DNA binding sites in the genome, instead of the better studied and more easily identified high-affinity binding sites.
"The sites that we have been considering good Hox sites don't provide the specificity that determines differences along the anterior-posterior axis of the animal body," Stern says. "High-affinity sites are not the sites that make animals animals." Stern, Mann, and their colleagues published their findings in the January 15, 2015, print edition of the journal Cell.
Hox genes produce proteins called transcription factors that help shape the identity of different segments of an animal's body by binding to DNA and switching on the appropriate genes. Fruit flies have eight different Hox genes; humans have more than 40. But strong similarities between the Hox proteins have mystified scientists for decades, Stern says.
"All of the Hox genes have very similar protein sequences in the domain that actually binds to DNA. Yet they must regulate different subsets of genes to make the head different from the thorax and different from the abdomen," he says. "It's not been at all clear how they do this."
Assorted experimental approaches failed to explain how different Hox proteins target unique sets of genes. The structures of various Hox genes suggested they all bind with high affinity to the same DNA sequence. And when Hox proteins are mixed with DNA in a test tube, they all tend to bind to the same sequence. Mann's discovery in 2009 that a protein cofactor often binds to DNA in tandem with Hox proteins offered a more complicated picture of Hox binding: When a Hox protein and its cofactor bind to adjacent pieces of DNA, they generate stable structures that recognize specific shapes in the DNA, and those new structures vary among Hox proteins. But in test tube experiments, Stern says, Hox proteins plus their cofactor still bind to similar sequences. "You get some specificity, but not a whole lot," he says. So the paradox persisted.
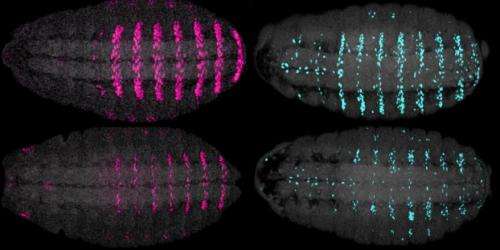
In his studies of genetic causes of species diversity, Stern has examined in detail the regulation of a gene called shavenbaby, which drives the production of a specific pattern of short, stout hairs, called trichomes, which are present on fruit fly larvae. Shavenbaby is switched on in stripes that run across certain segments of a fruit fly larva's body, suggesting it is likely under Hox control. So when Stern and Mann agreed to work together to figure out how Hox proteins achieve their specificity, shavenbaby seemed like a good place to start.
Crocker, a postdoctoral researcher in Stern's lab, demonstrated that the gene is in fact under the control of a Hox protein known as Ubx, showing that when Ubx is missing, no trichomes are produced in the Ubx domain. Conversely, by introducing Ubx to additional body segments, he could create flies with trichomes where they did not belong. He then showed that Ubx controls two specific enhancer regions that influence shavenbaby expression.
Ubx's influence over the shavenbaby enhancers seemed clear, yet when the scientists examined the DNA sequence of the enhancers, they could not find any regions that resembled classic Hox binding sites. So Mann's lab used biochemical experiments to search for physical evidence of Ubx binding. Scanning pieces of the enhancers one at a time, they found sites that Ubx bound to weakly, in the presence of the Hox cofactor. Each enhancer contained a cluster of these low-affinity binding sites. "These sequences didn't look like previously known Hox binding sites at all—none of them did," Stern says.
When Crocker created fruit flies with mutations in those sequences, shavenbaby expression and trichome production were impaired. One Ubx binding site was not enough to activate the gene, but when the cluster of binding sites was intact, enough Ubx bound to the enhancer to activate shavenbaby and turn on trichome production. The team had similar results with a shavenbaby enhancer from a different species of fruit fly. The scientists concluded that Hox proteins achieve specificity by binding to clusters of low-affinity sites in the genome. In fact, they say, many transcription factors likely bind to clusters of low-affinity sites, and their findings may apply broadly to gene regulation beyond the Hox family.
The findings explain why it has been difficult for scientists to predict where Hox proteins bind to DNA based on sequence data, Stern says. "The bioinformatic analysis has, to a first approximation, been useless for Hox genes. Now we know why. They're not binding to the sites everybody thought they were binding to—they're binding to these sites that don't even look like Hox binding sites."
"A lot of the field has been focused on the low-hanging fruit of these high-affinity sites, which are easier to identify computationally," Mann adds. "But we need to explore these low-affinity binding sites to really understand the specificity problem."
The study also underscores the importance of quantifying the effects of enhancer function, Stern says. When the team eliminated Hox binding sites in the shavenbaby enhancers one by one, the resulting changes in gene expression were subtle. "Every time I would do an experiment and analyze the data, it would look effectively the same," Crocker says. But by counting individual trichomes and developing other quantitative measures of gene expression, Crocker was able to provide convincing evidence of each site's importance.
The team went on to test how enhancer function was impacted when they exposed flies to less than optimal conditions: higher or lower temperatures, as well as a genetically reduced level of Ubx. Under those conditions, all of the Ubx binding sites were required to maintain normal shavenbaby activity. "This proves that these clusters of sites are there to provide robustness for the enhancer," Stern says. "At optimal conditions you don't really need all of the sites, but at anything other than perfectly optimal laboratory conditions you do. So in the wild, the flies need all of these sites."
More information: "Low Affinity Binding Site Clusters Confer Hox Specificity and Regulatory Robustness." Cell, DOI: dx.doi.org/10.1016/j.cell.2014.11.041
Journal information: Cell
Provided by Howard Hughes Medical Institute